| Entry | Database: PDB / ID: 5wht
|
|---|
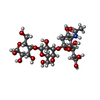
| Title | Crystal structure of 3'SL bound PltB |
|---|
 Components Components | Putative pertussis-like toxin subunit |
|---|
 Keywords Keywords | TOXIN / bacterial toxin / glycan |
|---|
| Function / homology |  Function and homology information Function and homology information
Bordetella pertussis toxin B, subunit 2/3, C-terminal / Pertussis toxin, subunit 2 and 3, C-terminal domain / OB fold (Dihydrolipoamide Acetyltransferase, E2P) - #110 / Enterotoxin / OB fold (Dihydrolipoamide Acetyltransferase, E2P) / Beta Barrel / Mainly BetaSimilarity search - Domain/homology 3'-sialyl-alpha-lactose / ACETATE ION / DI(HYDROXYETHYL)ETHER / N-acetyl-alpha-neuraminic acid / Pertussis-like toxin subunitSimilarity search - Component |
|---|
| Biological species |  Salmonella typhi (bacteria) Salmonella typhi (bacteria) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.932 Å MOLECULAR REPLACEMENT / Resolution: 1.932 Å |
|---|
 Authors Authors | Gao, X. / Galan, J.E. |
|---|
| Funding support |  United States, 1items United States, 1items | Organization | Grant number | Country |
|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | AI079022 |  United States United States |
|
|---|
 Citation Citation |  Journal: Nat Microbiol / Year: 2017 Journal: Nat Microbiol / Year: 2017
Title: Evolution of host adaptation in the Salmonella typhoid toxin.
Authors: Gao, X. / Deng, L. / Stack, G. / Yu, H. / Chen, X. / Naito-Matsui, Y. / Varki, A. / Galan, J.E. |
|---|
| History | | Deposition | Jul 18, 2017 | Deposition site: RCSB / Processing site: RCSB |
|---|
| Revision 1.0 | Oct 25, 2017 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Dec 6, 2017 | Group: Database references / Category: citation
Item: _citation.journal_volume / _citation.page_first / _citation.page_last |
|---|
| Revision 1.2 | Dec 11, 2019 | Group: Author supporting evidence / Data collection / Category: chem_comp / pdbx_audit_support
Item: _chem_comp.type / _pdbx_audit_support.funding_organization |
|---|
| Revision 2.0 | Jul 29, 2020 | Group: Atomic model / Data collection ...Atomic model / Data collection / Derived calculations / Structure summary
Category: atom_site / chem_comp ...atom_site / chem_comp / entity / entity_name_com / pdbx_branch_scheme / pdbx_chem_comp_identifier / pdbx_entity_branch / pdbx_entity_branch_descriptor / pdbx_entity_branch_link / pdbx_entity_branch_list / pdbx_entity_nonpoly / pdbx_molecule_features / pdbx_nonpoly_scheme / pdbx_struct_assembly_gen / struct_asym / struct_conn / struct_site / struct_site_gen
Item: _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ..._atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_asym_id / _atom_site.auth_atom_id / _atom_site.auth_comp_id / _atom_site.auth_seq_id / _atom_site.label_asym_id / _atom_site.label_atom_id / _atom_site.label_comp_id / _atom_site.label_entity_id / _atom_site.type_symbol / _chem_comp.name / _chem_comp.type / _entity.formula_weight / _entity.pdbx_description / _entity.pdbx_number_of_molecules / _entity.src_method / _entity.type / _pdbx_struct_assembly_gen.asym_id_list / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id
Description: Carbohydrate remediation / Provider: repository / Type: Remediation |
|---|
| Revision 2.1 | Oct 4, 2023 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Refinement description / Structure summary
Category: chem_comp / chem_comp_atom ...chem_comp / chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / struct_conn
Item: _chem_comp.pdbx_synonyms / _database_2.pdbx_DOI ..._chem_comp.pdbx_synonyms / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_conn.pdbx_leaving_atom_flag |
|---|
| Revision 2.2 | Nov 20, 2024 | Group: Structure summary / Category: pdbx_entry_details / pdbx_modification_feature |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Salmonella typhi (bacteria)
Salmonella typhi (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.932 Å
MOLECULAR REPLACEMENT / Resolution: 1.932 Å  Authors
Authors United States, 1items
United States, 1items  Citation
Citation Journal: Nat Microbiol / Year: 2017
Journal: Nat Microbiol / Year: 2017 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5wht.cif.gz
5wht.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5wht.ent.gz
pdb5wht.ent.gz PDB format
PDB format 5wht.json.gz
5wht.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/wh/5wht
https://data.pdbj.org/pub/pdb/validation_reports/wh/5wht ftp://data.pdbj.org/pub/pdb/validation_reports/wh/5wht
ftp://data.pdbj.org/pub/pdb/validation_reports/wh/5wht


 Links
Links Assembly
Assembly
 Components
Components Salmonella typhi (bacteria) / Gene: STY1891, t1107 / Production host:
Salmonella typhi (bacteria) / Gene: STY1891, t1107 / Production host: 





 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å
ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller













 PDBj
PDBj