+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5odm | |||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | NtiPr polyamide in complex with 5'CGATGTACTACG3 | |||||||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords KeywordsDNA / minor groove binders- hairpin polyamides | Function / homology | Chem-9T8 / 5-propan-2-yl-1,3-thiazole-4-carbaldehyde / GAMMA-AMINO-BUTANOIC ACID / 4-AMINO-(1-METHYLIMIDAZOLE)-2-CARBOXYLIC ACID / 4-AMINO-(1-METHYLPYRROLE)-2-CARBOXYLIC ACID / DNA / DNA (> 10) |  Function and homology information Function and homology informationBiological species |  Homo sapiens (human) Homo sapiens (human)Method | SOLUTION NMR / molecular dynamics / matrix relaxation |  Authors AuthorsPadroni, G. / Parkinson, J. / Burley, G.A. |  Citation Citation Journal: Nucleic Acids Res. / Year: 2018 Journal: Nucleic Acids Res. / Year: 2018Title: Structural basis of DNA duplex distortion induced by thiazole-containing hairpin polyamides. Authors: Padroni, G. / Parkinson, J.A. / Fox, K.R. / Burley, G.A. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5odm.cif.gz 5odm.cif.gz | 193.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5odm.ent.gz pdb5odm.ent.gz | 156.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5odm.json.gz 5odm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/od/5odm https://data.pdbj.org/pub/pdb/validation_reports/od/5odm ftp://data.pdbj.org/pub/pdb/validation_reports/od/5odm ftp://data.pdbj.org/pub/pdb/validation_reports/od/5odm | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5oczC 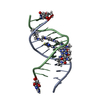 5odfC  5oe1C C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
-DNA chain , 1 types, 2 molecules AB
| #1: DNA chain | Mass: 3582.424 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.)  Homo sapiens (human) Homo sapiens (human) |
|---|
-Non-polymers , 5 types, 10 molecules 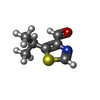
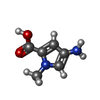

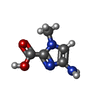




| #2: Chemical | ChemComp-9U2 / | ||||||
|---|---|---|---|---|---|---|---|
| #3: Chemical | ChemComp-PYB / #4: Chemical | ChemComp-ABU / | #5: Chemical | ChemComp-IMT / | #6: Chemical | ChemComp-9T8 / | |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||
| Sample conditions | Ionic strength: 100 phosphate mM / Label: Standard / pH: 7.4 / Pressure: 1 atm / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Bruker AVANCE / Manufacturer: Bruker / Model: AVANCE / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement |
| ||||||||||||||||||
| NMR ensemble | Conformer selection criteria: clustering / Conformers calculated total number: 2000 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller










 PDBj
PDBj











































 HSQC
HSQC