[English] 日本語
 Yorodumi
Yorodumi- PDB-5odh: Heterodisulfide reductase / [NiFe]-hydrogenase complex from Metha... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5odh | ||||||
|---|---|---|---|---|---|---|---|
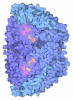
| Title | Heterodisulfide reductase / [NiFe]-hydrogenase complex from Methanothermococcus thermolithotrophicus soaked with heterodisulfide for 3.5 minutes | ||||||
 Components Components |
| ||||||
 Keywords Keywords | OXIDOREDUCTASE / Heterodisulfide reductase / [NiFe]-hydrogenase / FeS cluster / ferredoxin / CCG motif / methanogenesis / flavoprotein / flavin-based electron bifurcation / [2Fe-2S] cluster / macromolecular complex / anaerobic / thioredoxin / metabolism | ||||||
| Function / homology |  Function and homology information Function and homology informationdihydromethanophenazine:CoB-CoM heterodisulfide reductase / hydrogen dehydrogenase / hydrogen dehydrogenase activity / CoB--CoM heterodisulfide reductase activity / hydrogenase (acceptor) / methanogenesis / hydrogenase (acceptor) activity / Oxidoreductases; Acting on a sulfur group of donors / ferredoxin hydrogenase activity / nickel cation binding ...dihydromethanophenazine:CoB-CoM heterodisulfide reductase / hydrogen dehydrogenase / hydrogen dehydrogenase activity / CoB--CoM heterodisulfide reductase activity / hydrogenase (acceptor) / methanogenesis / hydrogenase (acceptor) activity / Oxidoreductases; Acting on a sulfur group of donors / ferredoxin hydrogenase activity / nickel cation binding / 2 iron, 2 sulfur cluster binding / 4 iron, 4 sulfur cluster binding / oxidoreductase activity / nucleotide binding / metal ion binding / plasma membrane Similarity search - Function | ||||||
| Biological species |  Methanothermococcus thermolithotrophicus DSM 2095 (archaea) Methanothermococcus thermolithotrophicus DSM 2095 (archaea) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | ||||||
 Authors Authors | Wagner, T. / Koch, J. / Ermler, U. / Shima, S. | ||||||
 Citation Citation |  Journal: Science / Year: 2017 Journal: Science / Year: 2017Title: Methanogenic heterodisulfide reductase (HdrABC-MvhAGD) uses two noncubane [4Fe-4S] clusters for reduction. Authors: Wagner, T. / Koch, J. / Ermler, U. / Shima, S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5odh.cif.gz 5odh.cif.gz | 815.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5odh.ent.gz pdb5odh.ent.gz | 655.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5odh.json.gz 5odh.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/od/5odh https://data.pdbj.org/pub/pdb/validation_reports/od/5odh ftp://data.pdbj.org/pub/pdb/validation_reports/od/5odh ftp://data.pdbj.org/pub/pdb/validation_reports/od/5odh | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5odcSC  5odiC  5odqC  5odrC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Heterodisulfide reductase, subunit ... , 3 types, 6 molecules AGBHCI
| #1: Protein | Mass: 71591.672 Da / Num. of mol.: 2 / Mutation: Wild-type / Source method: isolated from a natural source Source: (natural)  Methanothermococcus thermolithotrophicus DSM 2095 (archaea) Methanothermococcus thermolithotrophicus DSM 2095 (archaea)Cell line: / / Organ: / / Variant: / / Tissue: / / References: UniProt: A0A2D0TCB9*PLUS #2: Protein | Mass: 32075.318 Da / Num. of mol.: 2 / Mutation: Wild-type / Source method: isolated from a natural source Details: This subunit contains two pentacoordinated non cubane [4Fe-4S] clusters Source: (natural)  Methanothermococcus thermolithotrophicus DSM 2095 (archaea) Methanothermococcus thermolithotrophicus DSM 2095 (archaea)Cell line: / / Organ: / / Variant: / / Tissue: / References: UniProt: A0A2D0TCB4*PLUS, dihydromethanophenazine:CoB-CoM heterodisulfide reductase #3: Protein | Mass: 20483.650 Da / Num. of mol.: 2 / Mutation: Wild-type / Source method: isolated from a natural source Source: (natural)  Methanothermococcus thermolithotrophicus DSM 2095 (archaea) Methanothermococcus thermolithotrophicus DSM 2095 (archaea)Cell line: / / Organ: / / Variant: / / Tissue: / / References: UniProt: A0A2D0TC97*PLUS |
|---|
-Methyl-viologen reducing hydrogenase, subunit ... , 3 types, 6 molecules DJEKFL
| #4: Protein | Mass: 16057.495 Da / Num. of mol.: 2 / Mutation: Wild-type / Source method: isolated from a natural source Source: (natural)  Methanothermococcus thermolithotrophicus DSM 2095 (archaea) Methanothermococcus thermolithotrophicus DSM 2095 (archaea)Cell line: / / Organ: / / Variant: / / Tissue: / / References: UniProt: A0A2D0TC98*PLUS #5: Protein | Mass: 32511.432 Da / Num. of mol.: 2 / Mutation: Wild-type / Source method: isolated from a natural source Source: (natural)  Methanothermococcus thermolithotrophicus DSM 2095 (archaea) Methanothermococcus thermolithotrophicus DSM 2095 (archaea)Cell line: / / Organ: / / Variant: / / Tissue: / References: UniProt: A0A2D0TC99*PLUS, hydrogen dehydrogenase #6: Protein | Mass: 53129.602 Da / Num. of mol.: 2 / Mutation: Wild-type / Source method: isolated from a natural source Details: The C-terminus has been already post-translationally cleaved Source: (natural)  Methanothermococcus thermolithotrophicus DSM 2095 (archaea) Methanothermococcus thermolithotrophicus DSM 2095 (archaea)Cell line: / / Organ: / / Variant: / / Tissue: / References: UniProt: A0A2D0TCA6*PLUS, hydrogen dehydrogenase |
|---|
-Non-polymers , 13 types, 661 molecules 



















| #7: Chemical | ChemComp-SF4 / #8: Chemical | ChemComp-GOL / #9: Chemical | ChemComp-PE3 / #10: Chemical | #11: Chemical | ChemComp-9S8 / #12: Chemical | #13: Chemical | #14: Chemical | ChemComp-TRS / | #15: Chemical | #16: Chemical | #17: Chemical | #18: Chemical | ChemComp-TP7 / | #19: Water | ChemComp-HOH / | |
|---|
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.18 Å3/Da / Density % sol: 43.5 % Description: Black thick plate square shape of about 0.2-0.4 mm |
|---|---|
| Crystal grow | Temperature: 294 K / Method: vapor diffusion, sitting drop / pH: 8.5 Details: All crystallization was performed in an anaerobic chamber (95% N2/5% H2) with anoxic solution using the sitting drop method (96-well 2-drop MRC Crystallization Plates in polystyrene, ...Details: All crystallization was performed in an anaerobic chamber (95% N2/5% H2) with anoxic solution using the sitting drop method (96-well 2-drop MRC Crystallization Plates in polystyrene, Molecular Dimensions, Suffolk, UK). The crystallization reservoir contained 100 mM Tris/HCl, pH 8.5, 3% (v/v) DMSO, 30% (w/v) polyethylene glycol 4000, and 200 mM Na acetate trihydrate. Crystallization drop contained 1 ul HdrABC-MvhAGD at 25 mg/ml premixed with 2 mM FAD and 1 ul of precipitant. The crystals appeared after 1-2 weeks in this condition. The cryo-cool trapping experiment was performed as following: crystals were soaked for 3min30sec in the crystallization solution supplemented with 66 mM CoM-S-S-CoB. The crystal was cryo-protected by a soak in the crystallization solution supplemented by 30% glycerol and 10 mM CoM-S-S-CoB. |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X10SA / Wavelength: 0.97916 Å / Beamline: X10SA / Wavelength: 0.97916 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jun 23, 2016 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.97916 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→48.552 Å / Num. obs: 225651 / % possible obs: 99.8 % / Redundancy: 4.8 % / CC1/2: 0.992 / Rmerge(I) obs: 0.141 / Rpim(I) all: 0.073 / Net I/σ(I): 8.4 |
| Reflection shell | Resolution: 2.2→2.32 Å / Redundancy: 4.2 % / Rmerge(I) obs: 0.435 / Num. unique obs: 32668 / CC1/2: 0.451 / Rpim(I) all: 0.237 / % possible all: 99.5 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5ODC Resolution: 2.2→48.552 Å / SU ML: 0.33 / Cross valid method: THROUGHOUT / σ(F): 1.34 / Phase error: 25.47
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 40.4 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→48.552 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller








 PDBj
PDBj