+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5lxy | ||||||
|---|---|---|---|---|---|---|---|
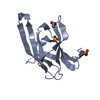
| Title | Structure of the minimal RBM7 - ZCCHC8 Complex | ||||||
 Components Components |
| ||||||
 Keywords Keywords | RNA BINDING PROTEIN / NEXT Complex RRM RBM7 ZCCHC8 | ||||||
| Function / homology |  Function and homology information Function and homology informationsnRNA catabolic process / TRAMP complex / co-transcriptional lncRNA 3' end processing, cleavage and polyadenylation pathway / snRNA binding / Nuclear RNA decay / mRNA 3'-end processing / regulation of alternative mRNA splicing, via spliceosome / pre-mRNA intronic binding / RNA processing / catalytic step 2 spliceosome ...snRNA catabolic process / TRAMP complex / co-transcriptional lncRNA 3' end processing, cleavage and polyadenylation pathway / snRNA binding / Nuclear RNA decay / mRNA 3'-end processing / regulation of alternative mRNA splicing, via spliceosome / pre-mRNA intronic binding / RNA processing / catalytic step 2 spliceosome / 14-3-3 protein binding / mRNA Splicing - Major Pathway / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / meiotic cell cycle / mRNA splicing, via spliceosome / single-stranded RNA binding / nuclear body / RNA binding / zinc ion binding / nucleoplasm / nucleus Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.85 Å MOLECULAR REPLACEMENT / Resolution: 2.85 Å | ||||||
 Authors Authors | Falk, S. / Finogenova, K. / Benda, C. / Conti, E. | ||||||
 Citation Citation |  Journal: Nat Commun / Year: 2016 Journal: Nat Commun / Year: 2016Title: Structure of the RBM7-ZCCHC8 core of the NEXT complex reveals connections to splicing factors. Authors: Falk, S. / Finogenova, K. / Melko, M. / Benda, C. / Lykke-Andersen, S. / Jensen, T.H. / Conti, E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5lxy.cif.gz 5lxy.cif.gz | 155.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5lxy.ent.gz pdb5lxy.ent.gz | 118.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5lxy.json.gz 5lxy.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lx/5lxy https://data.pdbj.org/pub/pdb/validation_reports/lx/5lxy ftp://data.pdbj.org/pub/pdb/validation_reports/lx/5lxy ftp://data.pdbj.org/pub/pdb/validation_reports/lx/5lxy | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5lxrSC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| 5 | 
| ||||||||
| 6 | 
| ||||||||
| 7 | 
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 10032.661 Da / Num. of mol.: 7 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: RBM7 / Production host: Homo sapiens (human) / Gene: RBM7 / Production host:  #2: Protein/peptide | Mass: 5097.909 Da / Num. of mol.: 7 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: ZCCHC8 / Production host: Homo sapiens (human) / Gene: ZCCHC8 / Production host:  #3: Chemical | ChemComp-BR / #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.9 Å3/Da / Density % sol: 57.3 % |
|---|---|
| Crystal grow | Temperature: 291.15 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: 0.1 M Bis-Tris-Propane 0.2 M NaBr 0.1 M Sodium malonate 20% (w/v) PEG3350 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X10SA / Wavelength: 1 Å / Beamline: X10SA / Wavelength: 1 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jun 26, 2015 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.85→71.79 Å / Num. obs: 24933 / % possible obs: 99.8 % / Redundancy: 3.2 % / CC1/2: 0.995 / Rmerge(I) obs: 0.061 / Net I/σ(I): 8.1 |
| Reflection shell | Resolution: 2.85→2.92 Å / CC1/2: 0.62 / % possible all: 99.7 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5LXR Resolution: 2.85→71.79 Å / Cor.coef. Fo:Fc: 0.866 / Cor.coef. Fo:Fc free: 0.829 / Cross valid method: THROUGHOUT / σ(F): 0 Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 244.62 Å2 / Biso min: 32.73 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.85→71.79 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.85→2.924 Å /
|
 Movie
Movie Controller
Controller












 PDBj
PDBj






