[English] 日本語
 Yorodumi
Yorodumi- PDB-5jlt: The crystal structure of the bacteriophage T4 MotA C-terminal dom... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5jlt | ||||||
|---|---|---|---|---|---|---|---|
| Title | The crystal structure of the bacteriophage T4 MotA C-terminal domain in complex with dsDNA reveals a novel protein-DNA recognition motif | ||||||
 Components Components |
| ||||||
 Keywords Keywords | VIRAL PROTEIN/DNA / MotA / dsDNA / "Double wing" / DNA binding motif / VIRAL PROTEIN-DNA complex | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus)synthetic construct (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.955 Å MOLECULAR REPLACEMENT / Resolution: 2.955 Å | ||||||
 Authors Authors | Cuypers, M.G. / Robertson, R.M. / Knipling, L. / Hinton, D.M. / White, S.W. | ||||||
 Citation Citation |  Journal: Nucleic Acids Res. / Year: 2018 Journal: Nucleic Acids Res. / Year: 2018Title: The phage T4 MotA transcription factor contains a novel DNA binding motif that specifically recognizes modified DNA. Authors: Cuypers, M.G. / Robertson, R.M. / Knipling, L. / Waddell, M.B. / Moon, K. / Hinton, D.M. / White, S.W. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5jlt.cif.gz 5jlt.cif.gz | 161.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5jlt.ent.gz pdb5jlt.ent.gz | 121.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5jlt.json.gz 5jlt.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  5jlt_validation.pdf.gz 5jlt_validation.pdf.gz | 483.8 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  5jlt_full_validation.pdf.gz 5jlt_full_validation.pdf.gz | 504.3 KB | Display | |
| Data in XML |  5jlt_validation.xml.gz 5jlt_validation.xml.gz | 27.2 KB | Display | |
| Data in CIF |  5jlt_validation.cif.gz 5jlt_validation.cif.gz | 39.2 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jl/5jlt https://data.pdbj.org/pub/pdb/validation_reports/jl/5jlt ftp://data.pdbj.org/pub/pdb/validation_reports/jl/5jlt ftp://data.pdbj.org/pub/pdb/validation_reports/jl/5jlt | HTTPS FTP |
-Related structure data
| Related structure data |  1kafS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 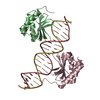
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14548.928 Da / Num. of mol.: 4 Source method: isolated from a genetically manipulated source Details: non-native amino acids from expression vector= EGDIHM. N-TERM RESIDUES (93-96) = ELLK. LINKER (97-104) = KRATRKAR. HTTP://WWW.UNIPROT.ORG/UNIPROT/P22915 Source: (gene. exp.)  Enterobacteria phage T4 (virus) / Gene: motA / Production host: Enterobacteria phage T4 (virus) / Gene: motA / Production host:  #2: DNA chain | Mass: 6710.366 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) #3: DNA chain | Mass: 6790.414 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.47 Å3/Da / Density % sol: 53.5 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: 23% PEG 8K, 0.1 M Na Acetate, 0.1 M NaCacodylate, pH 6.5, and 3% glycerol |
-Data collection
| Diffraction | Mean temperature: 100 K / Ambient temp details: cryostream |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 1 Å / Beamline: 22-ID / Wavelength: 1 Å |
| Detector | Type: MAR CCD 130 mm / Detector: CCD / Date: Jun 25, 2014 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.955→93.12 Å / Num. obs: 17332 / % possible obs: 100 % / Redundancy: 23.4 % / Biso Wilson estimate: 65.2 Å2 / CC1/2: 0.999 / Rmerge(I) obs: 0.167 / Net I/σ(I): 15.1 |
| Reflection shell | Resolution: 2.955→3.11 Å / Redundancy: 23.5 % / Mean I/σ(I) obs: 2 / % possible all: 100 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1KAF + DNA helix from COOT Resolution: 2.955→62.588 Å / Cross valid method: FREE R-VALUE / σ(F): 1.42 / Phase error: 35.46 Details: The settings of PHENIX.REFINE were tuned to use reference model restraints (PDB: 1KAF), NCS and the twin law K,H,-L.
| |||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | |||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 65.2 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.955→62.588 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller










 PDBj
PDBj







































