| Entry | Database: PDB / ID: 5epw
|
|---|
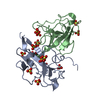
| Title | C-Terminal Domain Of Human Coronavirus Nl63 Nucleocapsid Protein |
|---|
 Components Components | Nucleoprotein |
|---|
 Keywords Keywords | VIRAL PROTEIN / NL63 coronavirus / N-protein / C-Terminal Domain |
|---|
| Function / homology |  Function and homology information Function and homology information
viral nucleocapsid / host cell endoplasmic reticulum-Golgi intermediate compartment / host cell cytoplasm / host cell Golgi apparatus / ribonucleoprotein complex / DNA binding / RNA bindingSimilarity search - Function Nucleocapsid protein, alphacoronavirus / Nucleocapsid protein, coronavirus / Nucleocapsid protein, C-terminal / Nucleocapsid protein, N-terminal / Nucleocapsid (N) protein, C-terminal domain, coronavirus / Nucleocapsid (N) protein, N-terminal domain, coronavirus / Coronavirus nucleocapsid / Coronavirus nucleocapsid (CoV N) protein N-terminal (NTD) domain profile. / Coronavirus nucleocapsid (CoV N) protein C-terminal (CTD) domain profile.Similarity search - Domain/homology |
|---|
| Biological species |  Human coronavirus NL63 Human coronavirus NL63 |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.5 Å MOLECULAR REPLACEMENT / Resolution: 1.5 Å |
|---|
 Authors Authors | Szelazek, B. / Kabala, W. / Kus, K. / Zdzalik, M. / Golik, P. / Florek, D. / Burmistrz, M. / Pyrc, K. / Dubin, G. |
|---|
| Funding support |  Poland, 1items Poland, 1items | Organization | Grant number | Country |
|---|
| National Science Center | UMO-2011/01/D/NZ1/01169 |  Poland Poland |
|
|---|
 Citation Citation |  Journal: J. Virol. / Year: 2017 Journal: J. Virol. / Year: 2017
Title: Structural Characterization of Human Coronavirus NL63 N Protein.
Authors: Szelazek, B. / Kabala, W. / Kus, K. / Zdzalik, M. / Twarda-Clapa, A. / Golik, P. / Burmistrz, M. / Florek, D. / Wladyka, B. / Pyrc, K. / Dubin, G. |
|---|
| History | | Deposition | Nov 12, 2015 | Deposition site: RCSB / Processing site: PDBE |
|---|
| Revision 1.0 | Feb 22, 2017 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Apr 5, 2017 | Group: Database references |
|---|
| Revision 1.2 | May 24, 2017 | Group: Database references |
|---|
| Revision 1.3 | May 8, 2024 | Group: Data collection / Database references / Category: chem_comp_atom / chem_comp_bond / database_2
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Human coronavirus NL63
Human coronavirus NL63 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.5 Å
MOLECULAR REPLACEMENT / Resolution: 1.5 Å  Authors
Authors Poland, 1items
Poland, 1items  Citation
Citation Journal: J. Virol. / Year: 2017
Journal: J. Virol. / Year: 2017 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5epw.cif.gz
5epw.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5epw.ent.gz
pdb5epw.ent.gz PDB format
PDB format 5epw.json.gz
5epw.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/ep/5epw
https://data.pdbj.org/pub/pdb/validation_reports/ep/5epw ftp://data.pdbj.org/pub/pdb/validation_reports/ep/5epw
ftp://data.pdbj.org/pub/pdb/validation_reports/ep/5epw Links
Links Assembly
Assembly
 Components
Components Human coronavirus NL63 / Gene: N, 6 / Production host:
Human coronavirus NL63 / Gene: N, 6 / Production host: 
 X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  Diamond
Diamond  / Beamline: I23 / Wavelength: 0.918409 Å
/ Beamline: I23 / Wavelength: 0.918409 Å Processing
Processing MOLECULAR REPLACEMENT / Resolution: 1.5→49.83 Å / Cor.coef. Fo:Fc: 0.968 / Cor.coef. Fo:Fc free: 0.96 / SU B: 1.337 / SU ML: 0.049 / Cross valid method: THROUGHOUT / ESU R: 0.066 / ESU R Free: 0.069 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
MOLECULAR REPLACEMENT / Resolution: 1.5→49.83 Å / Cor.coef. Fo:Fc: 0.968 / Cor.coef. Fo:Fc free: 0.96 / SU B: 1.337 / SU ML: 0.049 / Cross valid method: THROUGHOUT / ESU R: 0.066 / ESU R Free: 0.069 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller













 PDBj
PDBj