[English] 日本語
 Yorodumi
Yorodumi- PDB-5egb: Human PRDM9 allele-A ZnF Domain with Associated Recombination Hot... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5egb | ||||||
|---|---|---|---|---|---|---|---|
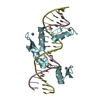
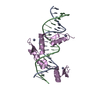
| Title | Human PRDM9 allele-A ZnF Domain with Associated Recombination Hotspot DNA Sequence II | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSCRIPTION/DNA / protein-DNA complex / zinc finger protein / TRANSCRIPTION-DNA complex | ||||||
| Function / homology |  Function and homology information Function and homology informationrecombination hotspot binding / positive regulation of reciprocal meiotic recombination / [histone H3]-lysine9 N-trimethyltransferase / male gamete generation / meiotic gene conversion / positive regulation of fertilization / [histone H4]-N-methyl-L-lysine20 N-methyltransferase / [histone H4]-lysine20 N-methyltransferase / histone H4K20 monomethyltransferase activity / female gamete generation ...recombination hotspot binding / positive regulation of reciprocal meiotic recombination / [histone H3]-lysine9 N-trimethyltransferase / male gamete generation / meiotic gene conversion / positive regulation of fertilization / [histone H4]-N-methyl-L-lysine20 N-methyltransferase / [histone H4]-lysine20 N-methyltransferase / histone H4K20 monomethyltransferase activity / female gamete generation / histone H3K9 trimethyltransferase activity / histone H4K20me methyltransferase activity / [histone H3]-lysine4 N-trimethyltransferase / [histone H3]-lysine36 N-trimethyltransferase / histone H3K36 trimethyltransferase activity / histone H3K4 trimethyltransferase activity / homologous chromosome pairing at meiosis / double-strand break repair involved in meiotic recombination / histone H3K36 methyltransferase activity / histone H3K4 methyltransferase activity / Transferases; Transferring one-carbon groups; Methyltransferases / PKMTs methylate histone lysines / Meiotic recombination / chromosome / regulation of gene expression / methylation / transcription cis-regulatory region binding / regulation of DNA-templated transcription / negative regulation of apoptotic process / protein homodimerization activity / zinc ion binding / nucleoplasm / nucleus Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.977 Å MOLECULAR REPLACEMENT / Resolution: 1.977 Å | ||||||
 Authors Authors | Patel, A. / Horton, J.R. / Wilson, G.G. / Zhang, X. / Cheng, X. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Genes Dev. / Year: 2016 Journal: Genes Dev. / Year: 2016Title: Structural basis for human PRDM9 action at recombination hot spots. Authors: Patel, A. / Horton, J.R. / Wilson, G.G. / Zhang, X. / Cheng, X. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5egb.cif.gz 5egb.cif.gz | 67.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5egb.ent.gz pdb5egb.ent.gz | 45.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5egb.json.gz 5egb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/eg/5egb https://data.pdbj.org/pub/pdb/validation_reports/eg/5egb ftp://data.pdbj.org/pub/pdb/validation_reports/eg/5egb ftp://data.pdbj.org/pub/pdb/validation_reports/eg/5egb | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 6489.186 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  Homo sapiens (human) Homo sapiens (human) | ||||
|---|---|---|---|---|---|
| #2: DNA chain | Mass: 6400.123 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  Homo sapiens (human) Homo sapiens (human) | ||||
| #3: Protein | Mass: 16982.342 Da / Num. of mol.: 1 / Fragment: ZnF8-12 (UNP residues 717-858) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: PRDM9, PFM6 / Variant: Allele-A / Plasmid: pGEX6p-1 / Production host: Homo sapiens (human) / Gene: PRDM9, PFM6 / Variant: Allele-A / Plasmid: pGEX6p-1 / Production host:  References: UniProt: Q9NQV7, histone-lysine N-methyltransferase | ||||
| #4: Chemical | ChemComp-ZN / #5: Water | ChemComp-HOH / | Has protein modification | N | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.12 Å3/Da / Density % sol: 60.61 % |
|---|---|
| Crystal grow | Temperature: 289 K / Method: vapor diffusion, hanging drop / pH: 5.5 Details: 0.055M Bis-Tris propane, 0.045M Citric Acid, 30% PEG3350 PH range: 5.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU MICROMAX-007 HF / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU MICROMAX-007 HF / Wavelength: 1.5418 Å |
| Detector | Type: RIGAKU SATURN 944+ / Detector: CCD / Date: May 5, 2015 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.97→29.64 Å / Num. obs: 22386 / % possible obs: 86.1 % / Redundancy: 7.7 % / Rmerge(I) obs: 0.071 / Net I/σ(I): 25.3 |
| Reflection shell | Resolution: 1.97→2.04 Å / Redundancy: 4.5 % / Rmerge(I) obs: 0.712 / Mean I/σ(I) obs: 1.9 / % possible all: 37.6 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: Zn SAD determined structure Resolution: 1.977→29.639 Å / SU ML: 0.19 / Cross valid method: FREE R-VALUE / σ(F): 1.33 / Phase error: 26.02 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.977→29.639 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj










































