| Entry | Database: PDB / ID: 4yhu
|
|---|
| Title | Yeast Prp3 C-terminal fragment 296-469 |
|---|
 Components Components | U4/U6 small nuclear ribonucleoprotein PRP3 |
|---|
 Keywords Keywords | RNA BINDING PROTEIN / Spliceosomal protein / DUF1115 / ferredoxin-like fold |
|---|
| Function / homology |  Function and homology information Function and homology information
U4/U6 snRNP / U4 snRNP / spliceosomal complex assembly / spliceosomal snRNP assembly / U4/U6 x U5 tri-snRNP complex / spliceosomal complex / mRNA splicing, via spliceosome / nucleusSimilarity search - Function Pre-mRNA-splicing factor 3 / U4/U6 small nuclear ribonucleoprotein Prp3 / pre-mRNA processing factor 3 domain / Small nuclear ribonucleoprotein Prp3, C-terminal domain / Small nuclear ribonucleoprotein Prp3, C-terminal domainSimilarity search - Domain/homology |
|---|
| Biological species |   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 2.7 Å MAD / Resolution: 2.7 Å |
|---|
 Authors Authors | Liu, S. / Wahl, M.C. |
|---|
| Funding support |  Germany, 1items Germany, 1items | Organization | Grant number | Country |
|---|
| German Research Foundation | SFB 740 |  Germany Germany |
|
|---|
 Citation Citation |  Journal: Elife / Year: 2015Title Journal: Elife / Year: 2015Title: A composite double-/single-stranded RNA-binding region in protein Prp3 supports tri-snRNP stability and splicing. Authors: Liu, S. / Mozaffari-Jovin, S. / Wollenhaupt, J. / Santos, K.F. / Theuser, M. / Dunin-Horkawicz, S. / Fabrizio, P. / Bujnicki, J.M. / Luhrmann, R. / Wahl, M.C. |
|---|
| History | | Deposition | Feb 27, 2015 | Deposition site: RCSB / Processing site: PDBE |
|---|
| Revision 1.0 | Jul 22, 2015 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Sep 30, 2015 | Group: Database references |
|---|
| Revision 2.0 | May 8, 2024 | Group: Advisory / Atomic model ...Advisory / Atomic model / Author supporting evidence / Data collection / Database references / Derived calculations
Category: atom_site / chem_comp_atom ...atom_site / chem_comp_atom / chem_comp_bond / database_2 / pdbx_audit_support / pdbx_nonpoly_scheme / pdbx_struct_conn_angle / pdbx_validate_close_contact / struct_conn / struct_site_gen
Item: _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ..._atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.occupancy / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_audit_support.funding_organization / _pdbx_nonpoly_scheme.auth_seq_num / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.value / _pdbx_validate_close_contact.auth_seq_id_1 / _pdbx_validate_close_contact.auth_seq_id_2 / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr1_symmetry / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_conn.ptnr2_symmetry / _struct_site_gen.auth_seq_id |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MAD / Resolution: 2.7 Å
MAD / Resolution: 2.7 Å  Authors
Authors Germany, 1items
Germany, 1items  Citation
Citation Journal: Elife / Year: 2015
Journal: Elife / Year: 2015 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4yhu.cif.gz
4yhu.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4yhu.ent.gz
pdb4yhu.ent.gz PDB format
PDB format 4yhu.json.gz
4yhu.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/yh/4yhu
https://data.pdbj.org/pub/pdb/validation_reports/yh/4yhu ftp://data.pdbj.org/pub/pdb/validation_reports/yh/4yhu
ftp://data.pdbj.org/pub/pdb/validation_reports/yh/4yhu Links
Links Assembly
Assembly


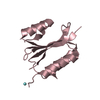
 Components
Components

 X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  BESSY
BESSY  / Beamline: 14.2 / Wavelength: 0.91841 Å
/ Beamline: 14.2 / Wavelength: 0.91841 Å Processing
Processing MAD / Resolution: 2.7→30 Å / Cor.coef. Fo:Fc: 0.946 / Cor.coef. Fo:Fc free: 0.926 / SU B: 25.673 / SU ML: 0.245 / Cross valid method: FREE R-VALUE / ESU R: 0.441 / ESU R Free: 0.306 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
MAD / Resolution: 2.7→30 Å / Cor.coef. Fo:Fc: 0.946 / Cor.coef. Fo:Fc free: 0.926 / SU B: 25.673 / SU ML: 0.245 / Cross valid method: FREE R-VALUE / ESU R: 0.441 / ESU R Free: 0.306 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller















 PDBj
PDBj



