+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4m9s | ||||||
|---|---|---|---|---|---|---|---|
| Title | crystal structure of CED-4 bound CED-3 fragment | ||||||
 Components Components |
| ||||||
 Keywords Keywords | APOPTOSIS / apoptosome / CED-3 | ||||||
| Function / homology |  Function and homology information Function and homology informationBH1 domain binding / regulation of development, heterochronic / positive regulation of apoptotic process involved in development / caspase complex / positive regulation of synapse pruning / peptidase activator activity involved in apoptotic process / caspase binding / positive regulation of protein processing / embryonic morphogenesis / apoptotic process involved in development ...BH1 domain binding / regulation of development, heterochronic / positive regulation of apoptotic process involved in development / caspase complex / positive regulation of synapse pruning / peptidase activator activity involved in apoptotic process / caspase binding / positive regulation of protein processing / embryonic morphogenesis / apoptotic process involved in development / cysteine-type endopeptidase activator activity / actin filament depolymerization / embryo development ending in birth or egg hatching / negative regulation of execution phase of apoptosis / regulation of cell size / cysteine-type endopeptidase activator activity involved in apoptotic process / muscle cell cellular homeostasis / BH3 domain binding / regulation of cell adhesion / endopeptidase activator activity / regulation of protein stability / ADP binding / defense response to Gram-negative bacterium / positive regulation of apoptotic process / apoptotic process / negative regulation of apoptotic process / perinuclear region of cytoplasm / magnesium ion binding / protein-containing complex / mitochondrion / ATP binding / membrane / identical protein binding / nucleus / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3.207 Å MOLECULAR REPLACEMENT / Resolution: 3.207 Å | ||||||
 Authors Authors | Huang, W.J. / Jinag, T.Y. / Choi, W.Y. / Wang, J.W. / Shi, Y.G. | ||||||
 Citation Citation |  Journal: Genes Dev. / Year: 2013 Journal: Genes Dev. / Year: 2013Title: Mechanistic insights into CED-4-mediated activation of CED-3. Authors: Huang, W. / Jiang, T. / Choi, W. / Qi, S. / Pang, Y. / Hu, Q. / Xu, Y. / Gong, X. / Jeffrey, P.D. / Wang, J. / Shi, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4m9s.cif.gz 4m9s.cif.gz | 425.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4m9s.ent.gz pdb4m9s.ent.gz | 350.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4m9s.json.gz 4m9s.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/m9/4m9s https://data.pdbj.org/pub/pdb/validation_reports/m9/4m9s ftp://data.pdbj.org/pub/pdb/validation_reports/m9/4m9s ftp://data.pdbj.org/pub/pdb/validation_reports/m9/4m9s | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4m9rC  4m9xC  4m9yC  4m9zC  3lqqS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 63797.398 Da / Num. of mol.: 4 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Protein/peptide | Mass: 1039.957 Da / Num. of mol.: 4 / Source method: obtained synthetically / Details: The peptide was chemically synthesized. #3: Chemical | ChemComp-MG / #4: Chemical | ChemComp-ATP / Has protein modification | Y | Sequence details | SEQUENCE OF THE PROTEIN WAS BASED ON ISOFORM A OF UNIPROT P30429 (CED4_CAEEL, IDENTIFIER | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.1 Å3/Da / Density % sol: 60.32 % Description: THE ENTRY CONTAINS FRIEDEL PAIRS IN F_PLUS/MINUS and I_PLUS/MINUS COLUMNS |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 0.6-0.8M sodium acetate, 0.1M HEPES (pH 7.5), 0.1M sodium fluoride , VAPOR DIFFUSION, HANGING DROP, temperature 291K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SPring-8 SPring-8  / Beamline: BL41XU / Wavelength: 0.97894 Å / Beamline: BL41XU / Wavelength: 0.97894 Å |
| Detector | Type: RAYONIX MX225HE / Detector: CCD / Date: Nov 12, 2012 |
| Radiation | Monochromator: Si 111 CHANNEL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.97894 Å / Relative weight: 1 |
| Reflection | Resolution: 3.2→50 Å / Num. obs: 56679 / % possible obs: 74.6 % / Observed criterion σ(F): 1 / Observed criterion σ(I): 1 |
| Reflection shell | Resolution: 3.2→3.31 Å / % possible all: 27.6 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3LQQ Resolution: 3.207→40.849 Å / SU ML: 0.56 / σ(F): 1.33 / Phase error: 38.79 / Stereochemistry target values: ML Details: THE ENTRY CONTAINS FRIEDEL PAIRS IN F_PLUS/MINUS and I_PLUS/MINUS COLUMNS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3.207→40.849 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller






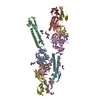




 PDBj
PDBj








