Entry Database : PDB / ID : 4l1lTitle Rat PKC C2 domain bound to CD Protein kinase C alpha type Keywords / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Rattus norvegicus (Norway rat)Method / / Resolution : 1.6 Å Authors Morales, K.M. / Yang, Y. / Long, Z. / Li, P. / Taylor, A.B. / Hart, P.J. / Igumenova, T.I. Journal : J.Am.Chem.Soc. / Year : 2013Title : Cd(2+) as a ca(2+) surrogate in protein-membrane interactions: isostructural but not isofunctional.Authors : Morales, K.A. / Yang, Y. / Long, Z. / Li, P. / Taylor, A.B. / Hart, P.J. / Igumenova, T.I. History Deposition Jun 3, 2013 Deposition site / Processing site Revision 1.0 Aug 28, 2013 Provider / Type Revision 1.1 Oct 23, 2013 Group Revision 1.2 Feb 28, 2024 Group / Database references / Derived calculationsCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_struct_conn_angle / struct_conn / struct_site Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_alt_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_alt_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.pdbx_ptnr1_label_alt_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SAD / Resolution: 1.6 Å
SAD / Resolution: 1.6 Å  Authors
Authors Citation
Citation Journal: J.Am.Chem.Soc. / Year: 2013
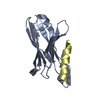
Journal: J.Am.Chem.Soc. / Year: 2013 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4l1l.cif.gz
4l1l.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4l1l.ent.gz
pdb4l1l.ent.gz PDB format
PDB format 4l1l.json.gz
4l1l.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/l1/4l1l
https://data.pdbj.org/pub/pdb/validation_reports/l1/4l1l ftp://data.pdbj.org/pub/pdb/validation_reports/l1/4l1l
ftp://data.pdbj.org/pub/pdb/validation_reports/l1/4l1l Links
Links Assembly
Assembly
 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU MICROMAX-007 HF / Wavelength: 1.54178 Å
ROTATING ANODE / Type: RIGAKU MICROMAX-007 HF / Wavelength: 1.54178 Å Processing
Processing SAD / Resolution: 1.6→43.816 Å / Isotropic thermal model: anisotropic / Cross valid method: THROUGHOUT / σ(F): 1.7 / Phase error: 17.9 / Stereochemistry target values: ML
SAD / Resolution: 1.6→43.816 Å / Isotropic thermal model: anisotropic / Cross valid method: THROUGHOUT / σ(F): 1.7 / Phase error: 17.9 / Stereochemistry target values: ML Movie
Movie Controller
Controller












 PDBj
PDBj













