[English] 日本語
 Yorodumi
Yorodumi- PDB-4i7s: T4 Lysozyme L99A/M102H with 3-trifluoromethyl-5-methyl pyrazole bound -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4i7s | ||||||
|---|---|---|---|---|---|---|---|
| Title | T4 Lysozyme L99A/M102H with 3-trifluoromethyl-5-methyl pyrazole bound | ||||||
 Components Components | Lysozyme | ||||||
 Keywords Keywords | HYDROLASE | ||||||
| Function / homology |  Function and homology information Function and homology informationviral release from host cell by cytolysis / peptidoglycan catabolic process / cell wall macromolecule catabolic process / lysozyme / lysozyme activity / host cell cytoplasm / defense response to bacterium Similarity search - Function | ||||||
| Biological species |  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.69 Å MOLECULAR REPLACEMENT / Resolution: 1.69 Å | ||||||
 Authors Authors | Merski, M. / Shoichet, B.K. | ||||||
 Citation Citation |  Journal: J.Med.Chem. / Year: 2013 Journal: J.Med.Chem. / Year: 2013Title: The impact of introducing a histidine into an apolar cavity site on docking and ligand recognition. Authors: Merski, M. / Shoichet, B.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4i7s.cif.gz 4i7s.cif.gz | 166 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4i7s.ent.gz pdb4i7s.ent.gz | 129.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4i7s.json.gz 4i7s.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/i7/4i7s https://data.pdbj.org/pub/pdb/validation_reports/i7/4i7s ftp://data.pdbj.org/pub/pdb/validation_reports/i7/4i7s ftp://data.pdbj.org/pub/pdb/validation_reports/i7/4i7s | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4i7jC  4i7kC  4i7lC  4i7mC  4i7nC  4i7oC  4i7pC  4i7qC  4i7rC  4i7tC  4e97S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The biological unit is a monomer. There are 2 biological units in the asymmetric unit (chains A and B) |
- Components
Components
-Protein , 1 types, 2 molecules AB
| #1: Protein | Mass: 21404.426 Da / Num. of mol.: 2 Mutation: T21C, S38D, L99A, M102H, E108V, S117V, T142C, N144D Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Enterobacteria phage T4 (virus) / Gene: E / Plasmid: pET-28 / Production host: Enterobacteria phage T4 (virus) / Gene: E / Plasmid: pET-28 / Production host:  |
|---|
-Non-polymers , 6 types, 370 molecules 
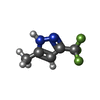


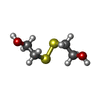






| #2: Chemical | | #3: Chemical | #4: Chemical | ChemComp-SO4 / #5: Chemical | #6: Chemical | ChemComp-HED / | #7: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.25 Å3/Da / Density % sol: 45.4 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 4.5 Details: 0.1 M sodium acetate, pH 4.5, 30% (w/v) PEG-6000, 0.3 M LiSO4, 3% (w/v) TMAO, 50 mM 2-mercaptoethanol, 50 mM 2-hydroxyethyl disulfide, vapor diffusion, hanging drop, temperature 277K |
-Data collection
| Diffraction | Mean temperature: 100 K | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ALS ALS  / Beamline: 8.3.1 / Wavelength: 1.116 Å / Beamline: 8.3.1 / Wavelength: 1.116 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Dec 9, 2010 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: two flat Si(111) crystals, mounted in a model DCM from Khozu Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.116 Å / Relative weight: 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.69→48.508 Å / Num. obs: 42023 / % possible obs: 98.8 % / Observed criterion σ(I): -3 / Biso Wilson estimate: 21.041 Å2 / Rmerge(I) obs: 0.056 / Net I/σ(I): 14.04 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 4E97 Resolution: 1.69→48.508 Å / Occupancy max: 1 / Occupancy min: 0.2 / FOM work R set: 0.8881 / SU ML: 0.43 / σ(F): 1.99 / Phase error: 18.66 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.95 Å / VDW probe radii: 1.2 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 41.939 Å2 / ksol: 0.372 e/Å3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 54.18 Å2 / Biso mean: 14.7495 Å2 / Biso min: 3.39 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.69→48.508 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Total num. of bins used: 12
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj











