+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4bzi | ||||||
|---|---|---|---|---|---|---|---|
| Title | The structure of the COPII coat assembled on membranes | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSPORT PROTEIN / SECRETION / TRAFFICKING / SEC23 / SEC24 / SAR1 | ||||||
| Function / homology |  Function and homology information Function and homology informationAntigen Presentation: Folding, assembly and peptide loading of class I MHC / Cargo concentration in the ER / : / positive regulation of ER to Golgi vesicle-mediated transport / COPII-mediated vesicle transport / mitochondria-associated endoplasmic reticulum membrane contact site / COPII-coated vesicle cargo loading / nuclear envelope organization / positive regulation of protein exit from endoplasmic reticulum / vesicle organization ...Antigen Presentation: Folding, assembly and peptide loading of class I MHC / Cargo concentration in the ER / : / positive regulation of ER to Golgi vesicle-mediated transport / COPII-mediated vesicle transport / mitochondria-associated endoplasmic reticulum membrane contact site / COPII-coated vesicle cargo loading / nuclear envelope organization / positive regulation of protein exit from endoplasmic reticulum / vesicle organization / COPII vesicle coat / membrane organization / signal sequence receptor activity / mitochondrial fission / fungal-type vacuole membrane / reticulophagy / mitochondrial membrane organization / endoplasmic reticulum exit site / endoplasmic reticulum to Golgi vesicle-mediated transport / vesicle-mediated transport / GTPase activator activity / SNARE binding / macroautophagy / ER to Golgi transport vesicle membrane / intracellular protein transport / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Golgi membrane / GTPase activity / endoplasmic reticulum membrane / GTP binding / endoplasmic reticulum / mitochondrion / zinc ion binding / metal ion binding Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | ELECTRON MICROSCOPY / electron tomography / cryo EM / Resolution: 23 Å | ||||||
 Authors Authors | Zanetti, G. / Prinz, S. / Daum, S. / Meister, A. / Schekman, R. / Bacia, K. / Briggs, J.A.G. | ||||||
 Citation Citation |  Journal: Elife / Year: 2013 Journal: Elife / Year: 2013Title: The structure of the COPII transport-vesicle coat assembled on membranes. Authors: Giulia Zanetti / Simone Prinz / Sebastian Daum / Annette Meister / Randy Schekman / Kirsten Bacia / John A G Briggs /  Abstract: Coat protein complex II (COPII) mediates formation of the membrane vesicles that export newly synthesised proteins from the endoplasmic reticulum. The inner COPII proteins bind to cargo and membrane, ...Coat protein complex II (COPII) mediates formation of the membrane vesicles that export newly synthesised proteins from the endoplasmic reticulum. The inner COPII proteins bind to cargo and membrane, linking them to the outer COPII components that form a cage around the vesicle. Regulated flexibility in coat architecture is essential for transport of a variety of differently sized cargoes, but structural data on the assembled coat has not been available. We have used cryo-electron tomography and subtomogram averaging to determine the structure of the complete, membrane-assembled COPII coat. We describe a novel arrangement of the outer coat and find that the inner coat can assemble into regular lattices. The data reveal how coat subunits interact with one another and with the membrane, suggesting how coordinated assembly of inner and outer coats can mediate and regulate packaging of vesicles ranging from small spheres to large tubular carriers. DOI:http://dx.doi.org/10.7554/eLife.00951.001. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4bzi.cif.gz 4bzi.cif.gz | 1017.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4bzi.ent.gz pdb4bzi.ent.gz | 789.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4bzi.json.gz 4bzi.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bz/4bzi https://data.pdbj.org/pub/pdb/validation_reports/bz/4bzi ftp://data.pdbj.org/pub/pdb/validation_reports/bz/4bzi ftp://data.pdbj.org/pub/pdb/validation_reports/bz/4bzi | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2428MC  2429C  2430C 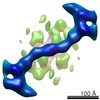 2431C  2432C  4bzjC  4bzkC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 | 
|
- Components
Components
-Protein , 4 types, 12 molecules ADGBJKELMFNO
| #1: Protein | Mass: 85463.242 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Plasmid: PTKY9 / Production host:  #2: Protein | Mass: 21472.564 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Plasmid: PTY40 / Production host:  #3: Protein | Mass: 103733.250 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Plasmid: PTKY9 / Production host:  #4: Protein | Mass: 103675.172 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Plasmid: PTKY9 / Production host:  |
|---|
-Non-polymers , 3 types, 12 molecules 




| #5: Chemical | ChemComp-ZN / #6: Chemical | #7: Chemical | |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: HELICAL ARRAY / 3D reconstruction method: electron tomography |
- Sample preparation
Sample preparation
| Component | Name: SEC23-23-SAR1 / Type: COMPLEX |
|---|---|
| Buffer solution | Name: MM HEPES, 50 MM KOAC, 1.2 MM MGCL2 / pH: 6.8 / Details: MM HEPES, 50 MM KOAC, 1.2 MM MGCL2 |
| Specimen | Conc.: 0.03 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Specimen support | Details: HOLEY CARBON |
| Vitrification | Instrument: HOMEMADE PLUNGER / Cryogen name: ETHANE / Details: LIQUID ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS / Date: Sep 18, 2012 |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 19500 X / Nominal defocus max: 3200 nm / Nominal defocus min: 2000 nm / Cs: 2.7 mm |
| Specimen holder | Tilt angle max: 60 ° / Tilt angle min: -60 ° |
| Image recording | Electron dose: 80 e/Å2 / Film or detector model: GATAN MULTISCAN |
| Image scans | Num. digital images: 26 |
- Processing
Processing
| EM software |
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Details: EACH TILTED IMAGE WITHIN TOMOGRAM | |||||||||||||||
| Symmetry | Point symmetry: C1 (asymmetric) | |||||||||||||||
| 3D reconstruction | Method: SUBTOMOGRAM ALIGNMENT AND AVERAGING / Resolution: 23 Å / Num. of particles: 15000 / Actual pixel size: 4.3 Å Details: SUBMISSION BASED ON EXPERIMENTAL DATA FROM EMDB EMD-2428. (DEPOSITION ID: 11856). Symmetry type: POINT | |||||||||||||||
| Atomic model building | Protocol: RIGID BODY FIT / Space: REAL / Target criteria: Cross-correlation coefficient / Details: METHOD--RIGID BODY REFINEMENT PROTOCOL--X-RAY | |||||||||||||||
| Atomic model building | PDB-ID: 1M2O Accession code: 1M2O / Source name: PDB / Type: experimental model | |||||||||||||||
| Refinement | Highest resolution: 23 Å | |||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 23 Å
|
 Movie
Movie Controller
Controller








 PDBj
PDBj











