Entry Database : PDB / ID : 3v43Title Crystal structure of MOZ Histone H3.1 Histone acetyltransferase KAT6A Keywords / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / / / Resolution : 1.47 Å Authors Qiu, Y. / Li, F. Journal : Genes Dev. / Year : 2012Title : Combinatorial readout of unmodified H3R2 and acetylated H3K14 by the tandem PHD finger of MOZ reveals a regulatory mechanism for HOXA9 transcriptionAuthors : Qiu, Y. / Liu, L. / Zhao, C. / Han, C. / Li, F. / Zhang, J. / Wang, Y. / Li, G. / Mei, Y. / Wu, M. / Wu, J. / Shi, Y. History Deposition Dec 14, 2011 Deposition site / Processing site Revision 1.0 Jun 27, 2012 Provider / Type Revision 1.1 Jul 17, 2013 Group Revision 1.2 Mar 20, 2024 Group / Database references / Derived calculationsCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_struct_conn_angle / struct_conn / struct_ref_seq_dif / struct_site Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_ref_seq_dif.details / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id Revision 1.3 Oct 30, 2024 Group / Category
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  SAD / Resolution: 1.47 Å
SAD / Resolution: 1.47 Å  Authors
Authors Citation
Citation Journal: Genes Dev. / Year: 2012
Journal: Genes Dev. / Year: 2012 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 3v43.cif.gz
3v43.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb3v43.ent.gz
pdb3v43.ent.gz PDB format
PDB format 3v43.json.gz
3v43.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/v4/3v43
https://data.pdbj.org/pub/pdb/validation_reports/v4/3v43 ftp://data.pdbj.org/pub/pdb/validation_reports/v4/3v43
ftp://data.pdbj.org/pub/pdb/validation_reports/v4/3v43 Links
Links Assembly
Assembly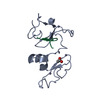
 Components
Components Homo sapiens (human) / Gene: KAT6A, MOZ, MYST3, RUNXBP2, ZNF220 / Plasmid: pET28a / Production host:
Homo sapiens (human) / Gene: KAT6A, MOZ, MYST3, RUNXBP2, ZNF220 / Plasmid: pET28a / Production host: 
 Homo sapiens (human) / References: UniProt: P68431
Homo sapiens (human) / References: UniProt: P68431 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SSRF
SSRF  / Beamline: BL17U / Wavelength: 0.9793 Å
/ Beamline: BL17U / Wavelength: 0.9793 Å SAD
SAD Processing
Processing SAD / Resolution: 1.47→19.13 Å / Cor.coef. Fo:Fc: 0.966 / Cor.coef. Fo:Fc free: 0.957 / Occupancy max: 1 / Occupancy min: 0 / SU B: 1.658 / SU ML: 0.03 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R Free: 0.055 / Stereochemistry target values: MAXIMUM LIKELIHOOD
SAD / Resolution: 1.47→19.13 Å / Cor.coef. Fo:Fc: 0.966 / Cor.coef. Fo:Fc free: 0.957 / Occupancy max: 1 / Occupancy min: 0 / SU B: 1.658 / SU ML: 0.03 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R Free: 0.055 / Stereochemistry target values: MAXIMUM LIKELIHOOD Movie
Movie Controller
Controller














 PDBj
PDBj


















