[English] 日本語
 Yorodumi
Yorodumi- PDB-2why: Crystal structure of the triscatecholate siderophore binding prot... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2why | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the triscatecholate siderophore binding protein FeuA from Bacillus subtilis complexed with Ferri-Bacillibactin | ||||||||||||
 Components Components |
| ||||||||||||
 Keywords Keywords | TRANSPORT PROTEIN / BACILLIBACTIN AND ENTEROBACTIN BINDING / TRISCATECHOLATE BINDING PROTEIN / IRON TRANSPORT / HIGH AFFINITY IRON IMPORT / IRON / MEMBRANE / PALMITATE / TRANSPORT / ABC-TYPE TRANSPORTER BINDING PROTEIN / SIDEROPHORE BINDING PROTEIN / LIPOPROTEIN / CELL MEMBRANE / ION TRANSPORT | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationiron coordination entity transport / outer membrane-bounded periplasmic space / membrane raft / plasma membrane / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |   | ||||||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | ||||||||||||
 Authors Authors | Peuckert, F. / Miethke, M. / Albrecht, A.G. / Essen, L.-O. / Marahiel, M.A. | ||||||||||||
 Citation Citation |  Journal: Angew.Chem.Int.Ed.Engl. / Year: 2009 Journal: Angew.Chem.Int.Ed.Engl. / Year: 2009Title: Structural Basis and Stereochemistry of Triscatecholate Siderophore Binding by Feua. Authors: Peuckert, F. / Miethke, M. / Albrecht, A.G. / Essen, L.-O. / Marahiel, M.A. | ||||||||||||
| History |
| ||||||||||||
| Remark 650 | HELIX DETERMINATION METHOD: AUTHOR PROVIDED. | ||||||||||||
| Remark 700 | SHEET DETERMINATION METHOD: AUTHOR PROVIDED. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2why.cif.gz 2why.cif.gz | 76.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2why.ent.gz pdb2why.ent.gz | 54.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2why.json.gz 2why.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/wh/2why https://data.pdbj.org/pub/pdb/validation_reports/wh/2why ftp://data.pdbj.org/pub/pdb/validation_reports/wh/2why ftp://data.pdbj.org/pub/pdb/validation_reports/wh/2why | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2wi8C  2phzS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 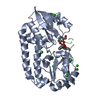
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| 2 | 
| ||||||||
| 3 | 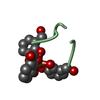
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 34746.660 Da / Num. of mol.: 1 / Fragment: RESIDUES 21-317 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   | ||||||
|---|---|---|---|---|---|---|---|
| #2: Protein/peptide | Mass: 900.796 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  | ||||||
| #3: Chemical | ChemComp-CL / #4: Chemical | ChemComp-FE / | #5: Water | ChemComp-HOH / | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.86 Å3/Da / Density % sol: 33.74 % / Description: NONE |
|---|---|
| Crystal grow | pH: 5.2 Details: PROTEIN WAS CRYSTALLIZED FROM 30% (V/V) PEG 600, 100 MM PHOSPHATE-CITRATE, PH 5.2; THEN SOAKED IN MOTHER LIQUOR CONTAINING 30% (V/V) GLYCEROL FOR CRYO PROTECTION. |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID29 / Wavelength: 0.9184 / Beamline: ID29 / Wavelength: 0.9184 |
| Detector | Type: ADSC CCD / Detector: CCD / Date: Nov 24, 2008 / Details: MIRRORS |
| Radiation | Monochromator: SI (111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9184 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→19.76 Å / Num. obs: 28022 / % possible obs: 99.2 % / Observed criterion σ(I): -3 / Redundancy: 3.7 % / Biso Wilson estimate: 24.416 Å2 / Rmerge(I) obs: 0.04 / Net I/σ(I): 20.1 |
| Reflection shell | Resolution: 1.7→1.79 Å / Redundancy: 3.7 % / Rmerge(I) obs: 0.52 / Mean I/σ(I) obs: 3.4 / % possible all: 99.5 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2PHZ Resolution: 1.7→19.51 Å / Cor.coef. Fo:Fc: 0.97 / Cor.coef. Fo:Fc free: 0.96 / SU B: 3.9 / SU ML: 0.07 / TLS residual ADP flag: LIKELY RESIDUAL / Cross valid method: THROUGHOUT / ESU R: 0.108 / ESU R Free: 0.103 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 23.303 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→19.51 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj








