[English] 日本語
 Yorodumi
Yorodumi- PDB-2w39: Glc(beta-1-3)Glc disaccharide in -1 and -2 sites of Laminarinase ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2w39 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
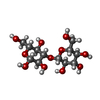
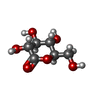
| Title | Glc(beta-1-3)Glc disaccharide in -1 and -2 sites of Laminarinase 16A from Phanerochaete chrysosporium | |||||||||
 Components Components | PUTATIVE LAMINARINASE | |||||||||
 Keywords Keywords | HYDROLASE / PHANEROCHAETE CHRYSOSPORIUM / WHITE ROT FUNGUS / GLYCOSYL HYDROLASE / GH7 / GH16 / LAM16A / LAMINARIN / FAMILY 16 / BETA-GLUCAN / BASIDIOMYCETE / BETA-GLUCANASE / PICHEA PASTORIS / LAMINARINASE / BETA SANDWICH / EXTRACELLULAR | |||||||||
| Function / homology |  Function and homology information Function and homology informationglucan catabolic process / hydrolase activity, hydrolyzing O-glycosyl compounds Similarity search - Function | |||||||||
| Biological species |  PHANEROCHAETE CHRYSOSPORIUM (fungus) PHANEROCHAETE CHRYSOSPORIUM (fungus) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.1 Å MOLECULAR REPLACEMENT / Resolution: 1.1 Å | |||||||||
 Authors Authors | Vasur, J. / Kawai, R. / Andersson, E. / Igarashi, K. / Sandgren, M. / Samejima, M. / Stahlberg, J. | |||||||||
 Citation Citation |  Journal: FEBS J. / Year: 2009 Journal: FEBS J. / Year: 2009Title: X-Ray Crystal Structures of Phanerochaete Chrysosporium Laminarinase 16A in Complex with Products from Lichenin and Laminarin Hydrolysis Authors: Vasur, J. / Kawai, R. / Andersson, E. / Igarashi, K. / Sandgren, M. / Samejima, M. / Stahlberg, J. | |||||||||
| History |
| |||||||||
| Remark 650 | HELIX DETERMINATION METHOD: AUTHOR PROVIDED. | |||||||||
| Remark 700 | SHEET DETERMINATION METHOD: AUTHOR PROVIDED. THE FOLLOWING SHEET RECORDS FOR CHAIN ID `A` HAVE ... SHEET DETERMINATION METHOD: AUTHOR PROVIDED. THE FOLLOWING SHEET RECORDS FOR CHAIN ID `A` HAVE BEEN GENERATED BY BETA-SPIDER, VERSION ALPHA 2.0 WITH AN ENERGY THRESHOLD OF -8.2 KCAL/MOL USING COULOMB ELECTROSTATICS USING 12-6 L-J VAN DER WAALS USING BETA-SPIDER RULE SETS. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2w39.cif.gz 2w39.cif.gz | 149.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2w39.ent.gz pdb2w39.ent.gz | 116.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2w39.json.gz 2w39.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/w3/2w39 https://data.pdbj.org/pub/pdb/validation_reports/w3/2w39 ftp://data.pdbj.org/pub/pdb/validation_reports/w3/2w39 ftp://data.pdbj.org/pub/pdb/validation_reports/w3/2w39 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2w52C  2cl2S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 31922.721 Da / Num. of mol.: 1 / Fragment: RESIDUES 21-318 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  PHANEROCHAETE CHRYSOSPORIUM (fungus) / Strain: K-3 / Plasmid: PPICZALPHAA / Production host: PHANEROCHAETE CHRYSOSPORIUM (fungus) / Strain: K-3 / Plasmid: PPICZALPHAA / Production host:  PICHIA PASTORIS (fungus) / Strain (production host): KM71H / References: UniProt: Q874E3, endo-1,3(4)-beta-glucanase PICHIA PASTORIS (fungus) / Strain (production host): KM71H / References: UniProt: Q874E3, endo-1,3(4)-beta-glucanase |
|---|---|
| #2: Polysaccharide | beta-D-glucopyranose-(1-3)-beta-D-glucopyranose / beta-laminaribiose |
| #3: Sugar | ChemComp-NAG / |
| #4: Sugar | ChemComp-LGC / |
| #5: Water | ChemComp-HOH / |
| Has protein modification | Y |
| Nonpolymer details | N-ACETYL GLUCOSAMIN |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.1 Å3/Da / Density % sol: 40.2 % / Description: NONE |
|---|---|
| Crystal grow | pH: 5 Details: 10 MG/ML LAMINARINASE 16A IN 25 % PEG 3350, 0.2 M AMMONIUM NITRATE AND 10 MM NAOAC PH 5, CO-CRYSTALIZED WITH 10 MM 4-O-BETA-D-GLUCOSYL-LAMINARIBIOSE (G4G3G) LIGAND FOR 6 MONTHS |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-1 / Wavelength: 0.934 / Beamline: ID14-1 / Wavelength: 0.934 |
| Detector | Type: ADSC CCD / Detector: CCD / Date: Dec 15, 2005 Details: DIAMOND (111), GE(220) SAGITALLY FOCUSING GE(220) AND A MULTILAYER |
| Radiation | Monochromator: DIAMOND (111), GE(220) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.934 Å / Relative weight: 1 |
| Reflection | Resolution: 1.1→29.55 Å / Num. obs: 98347 / % possible obs: 88.3 % / Observed criterion σ(I): 1.9 / Redundancy: 3.84 % / Biso Wilson estimate: 7 Å2 / Rmerge(I) obs: 0.06 / Net I/σ(I): 15.43 |
| Reflection shell | Resolution: 1.1→1.11 Å / Redundancy: 2.83 % / Rmerge(I) obs: 0.36 / Mean I/σ(I) obs: 2.3 / % possible all: 50.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2CL2 Resolution: 1.1→29.55 Å / Cor.coef. Fo:Fc: 0.974 / Cor.coef. Fo:Fc free: 0.97 / Cross valid method: THROUGHOUT / σ(F): 1.1 / ESU R: 0.033 / ESU R Free: 0.031 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK BULK SOLVENT | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 12.38 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.1→29.55 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj