[English] 日本語
 Yorodumi
Yorodumi- PDB-2oks: X-ray Structure of a DNA Repair Substrate Containing an Alkyl Int... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2oks | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | X-ray Structure of a DNA Repair Substrate Containing an Alkyl Interstrand Crosslink at 1.65 Resolution | ||||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / INTERSTRAND CROSSLINK / DNA DAMAGE | Function / homology | DNA |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  FOURIER SYNTHESIS / Resolution: 1.65 Å FOURIER SYNTHESIS / Resolution: 1.65 Å  Authors AuthorsSwenson, M.C. / Paranawithana, S.R. / Miller, P.S. / Kielkopf, C.L. |  Citation Citation Journal: Biochemistry / Year: 2007 Journal: Biochemistry / Year: 2007Title: Structure of a DNA repair substrate containing an alkyl interstrand cross-link at 1.65 a resolution. Authors: Swenson, M.C. / Paranawithana, S.R. / Miller, P.S. / Kielkopf, C.L. History |
Remark 600 | HETEROGEN THIS IS A N4C-ETHYL-N4C CROSSLINKED DNA. ONLY HALF OF THE ETHYL LINKER IS PRESENT IN THE ...HETEROGEN THIS IS A N4C-ETHYL-N4C CROSSLINKED DNA. ONLY HALF OF THE ETHYL LINKER IS PRESENT IN THE COORDINATES SINCE IT IS LOCATED ON THE TWO-FOLD AXES. | |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2oks.cif.gz 2oks.cif.gz | 19.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2oks.ent.gz pdb2oks.ent.gz | 10.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2oks.json.gz 2oks.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2oks_validation.pdf.gz 2oks_validation.pdf.gz | 374.4 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2oks_full_validation.pdf.gz 2oks_full_validation.pdf.gz | 374.3 KB | Display | |
| Data in XML |  2oks_validation.xml.gz 2oks_validation.xml.gz | 3.5 KB | Display | |
| Data in CIF |  2oks_validation.cif.gz 2oks_validation.cif.gz | 4.4 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ok/2oks https://data.pdbj.org/pub/pdb/validation_reports/ok/2oks ftp://data.pdbj.org/pub/pdb/validation_reports/ok/2oks ftp://data.pdbj.org/pub/pdb/validation_reports/ok/2oks | HTTPS FTP |
-Related structure data
| Related structure data |  1en8S S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 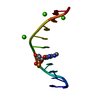
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 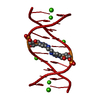
| ||||||||
| Unit cell |
| ||||||||
| Details | The biological assembly is a double-stranded DNA generated by the crystallographic two-fold axis |
- Components
Components
| #1: DNA chain | Mass: 3059.031 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: solid phase synthesis of DNA oligo, crosslink phosphoramidite synthesized separately | ||
|---|---|---|---|
| #2: Chemical | | #3: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.95 Å3/Da / Density % sol: 37.04 % | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 8 Details: The purified crosslinked DNA was dissolved in a buffer containing 22 mM ammonium acetate, 11 mM tris-hydroxymethyl-aminomethane (Tris)-HCl pH 8.0, and 11 mM calcium acetate. Crystals were ...Details: The purified crosslinked DNA was dissolved in a buffer containing 22 mM ammonium acetate, 11 mM tris-hydroxymethyl-aminomethane (Tris)-HCl pH 8.0, and 11 mM calcium acetate. Crystals were grown using the sitting drop vapor diffusion method at 277K, in which 3 uL of a reservoir solution containing 27% 2-3 methyl-pentadiol, 115 mM calcium acetate, and 10 mM Tris-HCl pH 8.0 was added to an equal volume of DNA solution, and the mixture was equilibrated against 700 uL of the reservoir solution. Crystals first appeared after two weeks. | ||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction | Mean temperature: 200 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.54 ROTATING ANODE / Type: RIGAKU / Wavelength: 1.54 |
| Detector | Type: RIGAKU RAXIS IV / Detector: IMAGE PLATE / Date: Aug 1, 2004 |
| Radiation | Monochromator: mirrors, graphite / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.54 Å / Relative weight: 1 |
| Reflection | Resolution: 1.65→40 Å / Num. all: 3043 / Num. obs: 2924 / % possible obs: 96.1 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 4.1 % / Biso Wilson estimate: 8 Å2 / Rsym value: 0.046 / Net I/σ(I): 38.1 |
| Reflection shell | Resolution: 1.65→1.71 Å / Redundancy: 2.56 % / Mean I/σ(I) obs: 25.7 / Num. unique all: 239 / Rsym value: 0.047 / % possible all: 79.4 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  FOURIER SYNTHESIS FOURIER SYNTHESISStarting model: PDB ENTRY 1EN8 Resolution: 1.65→18.89 Å / Rfactor Rfree error: 0.016 / Data cutoff high absF: 992530.75 / Data cutoff low absF: 0 / Isotropic thermal model: isotropic individual / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 25.6312 Å2 / ksol: 0.353499 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 12.8 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.65→18.89 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.65→1.75 Å / Rfactor Rfree error: 0.044 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller





 PDBj
PDBj




