+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2m5n | ||||||
|---|---|---|---|---|---|---|---|
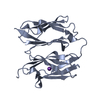
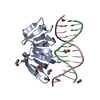
| Title | Atomic-resolution structure of a cross-beta protofilament | ||||||
 Components Components | Transthyretin | ||||||
 Keywords Keywords | PROTEIN FIBRIL / Amyloid fibril / Cross-beta structure | ||||||
| Function / homology |  Function and homology information Function and homology informationDefective visual phototransduction due to STRA6 loss of function / negative regulation of glomerular filtration / The canonical retinoid cycle in rods (twilight vision) / purine nucleobase metabolic process / hormone binding / Non-integrin membrane-ECM interactions / phototransduction, visible light / molecular sequestering activity / retinoid metabolic process / Retinoid metabolism and transport ...Defective visual phototransduction due to STRA6 loss of function / negative regulation of glomerular filtration / The canonical retinoid cycle in rods (twilight vision) / purine nucleobase metabolic process / hormone binding / Non-integrin membrane-ECM interactions / phototransduction, visible light / molecular sequestering activity / retinoid metabolic process / Retinoid metabolism and transport / hormone activity / azurophil granule lumen / Amyloid fiber formation / Neutrophil degranulation / protein-containing complex binding / protein-containing complex / : / extracellular exosome / extracellular region / identical protein binding Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | SOLID-STATE NMR / simulated annealing | ||||||
| Model details | lowest energy, model1 | ||||||
 Authors Authors | Fitzpatrick, A.W.P. / Debelouchina, G.T. / Bayro, M.J. / Clare, D.K. / Caporini, M.A. / Bajaj, V.S. / Jaroniec, C.P. / Wang, L. / Ladizhansky, V. / Muller, S. ...Fitzpatrick, A.W.P. / Debelouchina, G.T. / Bayro, M.J. / Clare, D.K. / Caporini, M.A. / Bajaj, V.S. / Jaroniec, C.P. / Wang, L. / Ladizhansky, V. / Muller, S. / MacPhee, C.E. / Waudby, C.A. / Mott, H.R. / de Simone, A. / Knowles, T.P.J. / Saibil, H.R. / Vendruscolo, M. / Orlova, E.V. / Griffin, R.G. / Dobson, C.M. | ||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2013 Journal: Proc Natl Acad Sci U S A / Year: 2013Title: Atomic structure and hierarchical assembly of a cross-β amyloid fibril. Authors: Anthony W P Fitzpatrick / Galia T Debelouchina / Marvin J Bayro / Daniel K Clare / Marc A Caporini / Vikram S Bajaj / Christopher P Jaroniec / Luchun Wang / Vladimir Ladizhansky / Shirley A ...Authors: Anthony W P Fitzpatrick / Galia T Debelouchina / Marvin J Bayro / Daniel K Clare / Marc A Caporini / Vikram S Bajaj / Christopher P Jaroniec / Luchun Wang / Vladimir Ladizhansky / Shirley A Müller / Cait E MacPhee / Christopher A Waudby / Helen R Mott / Alfonso De Simone / Tuomas P J Knowles / Helen R Saibil / Michele Vendruscolo / Elena V Orlova / Robert G Griffin / Christopher M Dobson /  Abstract: The cross-β amyloid form of peptides and proteins represents an archetypal and widely accessible structure consisting of ordered arrays of β-sheet filaments. These complex aggregates have ...The cross-β amyloid form of peptides and proteins represents an archetypal and widely accessible structure consisting of ordered arrays of β-sheet filaments. These complex aggregates have remarkable chemical and physical properties, and the conversion of normally soluble functional forms of proteins into amyloid structures is linked to many debilitating human diseases, including several common forms of age-related dementia. Despite their importance, however, cross-β amyloid fibrils have proved to be recalcitrant to detailed structural analysis. By combining structural constraints from a series of experimental techniques spanning five orders of magnitude in length scale--including magic angle spinning nuclear magnetic resonance spectroscopy, X-ray fiber diffraction, cryoelectron microscopy, scanning transmission electron microscopy, and atomic force microscopy--we report the atomic-resolution (0.5 Å) structures of three amyloid polymorphs formed by an 11-residue peptide. These structures reveal the details of the packing interactions by which the constituent β-strands are assembled hierarchically into protofilaments, filaments, and mature fibrils. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2m5n.cif.gz 2m5n.cif.gz | 1 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2m5n.ent.gz pdb2m5n.ent.gz | 884.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2m5n.json.gz 2m5n.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/m5/2m5n https://data.pdbj.org/pub/pdb/validation_reports/m5/2m5n ftp://data.pdbj.org/pub/pdb/validation_reports/m5/2m5n ftp://data.pdbj.org/pub/pdb/validation_reports/m5/2m5n | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2323C  2324C  5590C  2m5kC  2m5mC  3zpkC C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 1198.366 Da / Num. of mol.: 16 / Fragment: UNP residues 125-135 / Source method: obtained synthetically / Source: (synth.)  Homo sapiens (human) / References: UniProt: P02766 Homo sapiens (human) / References: UniProt: P02766 |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLID-STATE NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 15 mg/mL [U-100% 13C; U-100% 15N] TTR(105-115), 10% acetonitrile/water solution Solvent system: 10% acetonitrile/water solution |
|---|---|
| Sample | Conc.: 15 mg/mL / Component: TTR(105-115)-1 / Isotopic labeling: [U-100% 13C; U-100% 15N] |
| Sample conditions | pH: 2 / Pressure: ambient / Temperature units: K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 1 | |||||||||
| NMR representative | Selection criteria: lowest energy | |||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations Conformers calculated total number: 100 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller







 PDBj
PDBj




