+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2k6t | ||||||
|---|---|---|---|---|---|---|---|
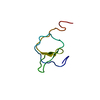
| Title | Solution structure of the relaxin-like factor | ||||||
 Components Components |
| ||||||
 Keywords Keywords | HORMONE / protein / Cleavage on pair of basic residues / Disease mutation / Polymorphism / Secreted | ||||||
| Function / homology |  Function and homology information Function and homology informationRelaxin receptors / positive regulation of wound healing / positive regulation of epithelial cell migration / insulin receptor binding / hormone activity / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / cell-cell signaling / protease binding / spermatogenesis / G alpha (s) signalling events ...Relaxin receptors / positive regulation of wound healing / positive regulation of epithelial cell migration / insulin receptor binding / hormone activity / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / cell-cell signaling / protease binding / spermatogenesis / G alpha (s) signalling events / signaling receptor binding / : / extracellular region Similarity search - Function | ||||||
| Method | SOLUTION NMR / torsion angle dynamics, simulated annealing | ||||||
 Authors Authors | Bullesbach, E.E. / Hass, M.A.S. / Jensen, M.R. / Hansen, D.F. / Kristensen, S.M. / Schwabe, C. / Led, J.J. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2008 Journal: Biochemistry / Year: 2008Title: Solution structure of a conformationally restricted fully active derivative of the human relaxin-like factor Authors: Bullesbach, E.E. / Hass, M.A.S. / Jensen, M.R. / Hansen, D.F. / Kristensen, S.M. / Schwabe, C. / Led, J.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2k6t.cif.gz 2k6t.cif.gz | 338.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2k6t.ent.gz pdb2k6t.ent.gz | 280.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2k6t.json.gz 2k6t.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/k6/2k6t https://data.pdbj.org/pub/pdb/validation_reports/k6/2k6t ftp://data.pdbj.org/pub/pdb/validation_reports/k6/2k6t ftp://data.pdbj.org/pub/pdb/validation_reports/k6/2k6t | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 2778.189 Da / Num. of mol.: 1 / Fragment: UNP residues 106-131 / Source method: obtained synthetically / Details: This sequence occurs naturally in humans. / References: UniProt: P51460 |
|---|---|
| #2: Protein/peptide | Mass: 3528.118 Da / Num. of mol.: 1 / Fragment: UNP residues 25-55 / Source method: obtained synthetically / Details: This sequence occurs naturally in humans. / References: UniProt: P51460 |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NMR details | Text: structure was determined using homonuclear NOE |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||||||||||||||||||||||
| Sample conditions |
|
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: torsion angle dynamics, simulated annealing / Software ordinal: 1 | ||||||||||||||||||||||||||||||||||||||||
| NMR constraints | NOE constraints total: 1154 / NOE intraresidue total count: 506 / NOE long range total count: 182 / NOE medium range total count: 208 / NOE sequential total count: 258 | ||||||||||||||||||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 100 / Conformers submitted total number: 20 / Maximum lower distance constraint violation: 0 Å / Maximum upper distance constraint violation: 6 Å / Representative conformer: 1 | ||||||||||||||||||||||||||||||||||||||||
| NMR ensemble rms | Distance rms dev: 0.053 Å / Distance rms dev error: 0.0015 Å |
 Movie
Movie Controller
Controller












 PDBj
PDBj




 HSQC
HSQC