[English] 日本語
 Yorodumi
Yorodumi- PDB-2imm: Refined crystal structure of a recombinant immunoglobulin domain ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2imm | ||||||
|---|---|---|---|---|---|---|---|
| Title | Refined crystal structure of a recombinant immunoglobulin domain and a complementarity-determining region 1-grafted mutant | ||||||
 Components Components | IGA-KAPPA MCPC603 FV (LIGHT CHAIN) | ||||||
 Keywords Keywords | IMMUNOGLOBULIN | ||||||
| Function / homology | Immunoglobulins / Immunoglobulin-like / Sandwich / Mainly Beta / ACETATE ION / :  Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2 Å X-RAY DIFFRACTION / Resolution: 2 Å | ||||||
 Authors Authors | Steipe, B. / Huber, R. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1992 Journal: J.Mol.Biol. / Year: 1992Title: Refined crystal structure of a recombinant immunoglobulin domain and a complementarity-determining region 1-grafted mutant. Authors: Steipe, B. / Pluckthun, A. / Huber, R. #1:  Journal: J.Mol.Biol. / Year: 1990 Journal: J.Mol.Biol. / Year: 1990Title: Crystallization and Preliminary X-Ray Studies of the VL Domain of the Antibody Mc/Pc603 Produced in Escherichia Coli Authors: Glockshuber, R. / Steipe, B. / Huber, R. / Pluckthun, A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2imm.cif.gz 2imm.cif.gz | 38 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2imm.ent.gz pdb2imm.ent.gz | 25.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2imm.json.gz 2imm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2imm_validation.pdf.gz 2imm_validation.pdf.gz | 385.1 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2imm_full_validation.pdf.gz 2imm_full_validation.pdf.gz | 385.2 KB | Display | |
| Data in XML |  2imm_validation.xml.gz 2imm_validation.xml.gz | 3.9 KB | Display | |
| Data in CIF |  2imm_validation.cif.gz 2imm_validation.cif.gz | 6.2 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/im/2imm https://data.pdbj.org/pub/pdb/validation_reports/im/2imm ftp://data.pdbj.org/pub/pdb/validation_reports/im/2imm ftp://data.pdbj.org/pub/pdb/validation_reports/im/2imm | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 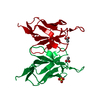
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Atom site foot note | 1: RESIDUES PRO 8 AND PRO 95 ARE CIS-PROLINES. 2: RESIDUES LYS 103 AND ARG 108 ARE PARTIALLY DISORDERED IN THE ELECTRON DENSITY. | ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Antibody | Mass: 12341.727 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  |
|---|---|
| #2: Chemical | ChemComp-SO4 / |
| #3: Chemical | ChemComp-ACT / |
| #4: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.26 Å3/Da / Density % sol: 62.3 % | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS pH: 4 / Method: unknown | |||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Reflection | *PLUS Highest resolution: 1.9 Å / Num. obs: 11262 / Num. measured all: 96963 / Rmerge(I) obs: 0.086 |
|---|
- Processing
Processing
| Software | Name: EREF / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Rfactor Rwork: 0.149 / Highest resolution: 2 Å Details: RESIDUES LYS 103 AND ARG 108 ARE PARTIALLY DISORDERED IN THE ELECTRON DENSITY | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 2 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2 Å / Rfactor obs: 0.149 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: o_angle_d / Dev ideal: 2.23 |
 Movie
Movie Controller
Controller












 PDBj
PDBj






