+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2erw | ||||||
|---|---|---|---|---|---|---|---|
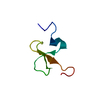
| Title | Crystal Structure of Infestin 4, a factor XIIa inhibitor | ||||||
 Components Components | serine protease inhibitor infestin | ||||||
 Keywords Keywords | BLOOD CLOTTING / HYDROLASE INHIBITOR / Kazal type domain | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Triatoma infestans (insect) Triatoma infestans (insect) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.4 Å MOLECULAR REPLACEMENT / Resolution: 1.4 Å | ||||||
 Authors Authors | Campos, I.T.N. / Tanaka, A.S. / Barbosa, J.A.R.G. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 2012 Journal: Acta Crystallogr.,Sect.D / Year: 2012Title: The Kazal-type inhibitors infestins 1 and 4 differ in specificity but are similar in three-dimensional structure. Authors: Campos, I.T. / Souza, T.A. / Torquato, R.J. / De Marco, R. / Tanaka-Azevedo, A.M. / Tanaka, A.S. / Barbosa, J.A.R.G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2erw.cif.gz 2erw.cif.gz | 24.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2erw.ent.gz pdb2erw.ent.gz | 15.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2erw.json.gz 2erw.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2erw_validation.pdf.gz 2erw_validation.pdf.gz | 398.2 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2erw_full_validation.pdf.gz 2erw_full_validation.pdf.gz | 399.5 KB | Display | |
| Data in XML |  2erw_validation.xml.gz 2erw_validation.xml.gz | 3.2 KB | Display | |
| Data in CIF |  2erw_validation.cif.gz 2erw_validation.cif.gz | 4.3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/er/2erw https://data.pdbj.org/pub/pdb/validation_reports/er/2erw ftp://data.pdbj.org/pub/pdb/validation_reports/er/2erw ftp://data.pdbj.org/pub/pdb/validation_reports/er/2erw | HTTPS FTP |
-Related structure data
| Related structure data |  2f3cC  1tbrS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Details | Asymmetric unit content is the biological assembly. |
- Components
Components
| #1: Protein | Mass: 6198.125 Da / Num. of mol.: 1 / Fragment: residues 167-222 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Triatoma infestans (insect) / Plasmid: pPIC9 / Production host: Triatoma infestans (insect) / Plasmid: pPIC9 / Production host:  Pichia pastoris (fungus) / Strain (production host): GS115 / References: UniProt: Q95P16 Pichia pastoris (fungus) / Strain (production host): GS115 / References: UniProt: Q95P16 |
|---|---|
| #2: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.68 Å3/Da / Density % sol: 54.03 % |
|---|---|
| Crystal grow | Temperature: 303 K / Method: vapor diffusion, sitting drop / pH: 5.6 Details: 30% PEG 4000, 0.1 M ammonium acetate, 0.1 M sodium cacodylate, pH 5.6, VAPOR DIFFUSION, SITTING DROP, temperature 303K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  LNLS LNLS  / Beamline: D03B-MX1 / Wavelength: 1.427 Å / Beamline: D03B-MX1 / Wavelength: 1.427 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Sep 9, 2004 |
| Radiation | Monochromator: Si CRYSTAL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.427 Å / Relative weight: 1 |
| Reflection | Resolution: 1.4→40 Å / Num. all: 13756 / Num. obs: 13618 / % possible obs: 99 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 7.1 % / Biso Wilson estimate: 15.315 Å2 / Rmerge(I) obs: 0.037 / Net I/σ(I): 52.5 |
| Reflection shell | Resolution: 1.4→1.45 Å / Redundancy: 5.3 % / Rmerge(I) obs: 0.367 / Mean I/σ(I) obs: 5.7 / Num. unique all: 1229 / % possible all: 91 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1TBR Resolution: 1.4→35.71 Å / Cor.coef. Fo:Fc: 0.961 / Cor.coef. Fo:Fc free: 0.962 / SU B: 1.712 / SU ML: 0.034 / TLS residual ADP flag: LIKELY RESIDUAL / Isotropic thermal model: Isotropic / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.055 / ESU R Free: 0.053 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: SIMULATED ANNEALING WAS PERFORMED WITH THE CNS SOFTWARE IN THE FIRST CYCLE OF REFINEMENT. TLS APPLIED USING THE PROTEIN MODEL AS ONE GROUP. Close contacts in remark 500 are related to the ...Details: SIMULATED ANNEALING WAS PERFORMED WITH THE CNS SOFTWARE IN THE FIRST CYCLE OF REFINEMENT. TLS APPLIED USING THE PROTEIN MODEL AS ONE GROUP. Close contacts in remark 500 are related to the alternate conformations of the CYS 6 region close to the N-terminus and CYS 31. The alternate conformation of the N-terminus is not modelled due to the weak electron density.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 24.051 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.4→35.71 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.403→1.439 Å / Total num. of bins used: 20
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 12.833 Å / Origin y: 7.551 Å / Origin z: 24.913 Å
|
 Movie
Movie Controller
Controller









 PDBj
PDBj

