+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2ds8 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of the ZBD-XB complex | ||||||
 Components Components |
| ||||||
 Keywords Keywords | METAL BINDING PROTEIN / PROTEIN BINDING / Protein-peptide complex | ||||||
| Function / homology |  Function and homology information Function and homology informationHslUV protease complex / endopeptidase Clp complex / ATP-dependent peptidase activity / protein unfolding / : / ATP-dependent protein folding chaperone / disordered domain specific binding / : / protease binding / protein dimerization activity ...HslUV protease complex / endopeptidase Clp complex / ATP-dependent peptidase activity / protein unfolding / : / ATP-dependent protein folding chaperone / disordered domain specific binding / : / protease binding / protein dimerization activity / cell division / ATP hydrolysis activity / zinc ion binding / ATP binding / identical protein binding / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.6 Å MOLECULAR REPLACEMENT / Resolution: 1.6 Å | ||||||
 Authors Authors | Park, E.Y. / Lee, B.G. / Hong, S.B. / Kim, H.W. / Song, H.K. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2007 Journal: J.Mol.Biol. / Year: 2007Title: Structural Basis of SspB-tail Recognition by the Zinc Binding Domain of ClpX. Authors: Park, E.Y. / Lee, B.G. / Hong, S.B. / Kim, H.W. / Jeon, H. / Song, H.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2ds8.cif.gz 2ds8.cif.gz | 33.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2ds8.ent.gz pdb2ds8.ent.gz | 22 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2ds8.json.gz 2ds8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ds/2ds8 https://data.pdbj.org/pub/pdb/validation_reports/ds/2ds8 ftp://data.pdbj.org/pub/pdb/validation_reports/ds/2ds8 ftp://data.pdbj.org/pub/pdb/validation_reports/ds/2ds8 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 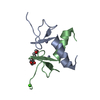 2ds5SC  2ds6C 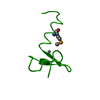 2ds7C S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 5758.631 Da / Num. of mol.: 2 / Fragment: Zinc binding domain(ZBD) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Protein/peptide | Mass: 855.079 Da / Num. of mol.: 2 / Source method: obtained synthetically Details: XB peptide (APALRVVK) is synthesized. Except the first alanine, the sequence occurs naturally in E. coli SspB #3: Chemical | #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.633 Å3/Da / Density % sol: 24.68 % |
|---|---|
| Crystal grow | Temperature: 295 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: 1.6M tri-sodium citrate, pH 6.5, 10-fold molar excess of XB peptide addition, VAPOR DIFFUSION, HANGING DROP, temperature 295K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Photon Factory Photon Factory  / Beamline: AR-NW12A / Wavelength: 1 Å / Beamline: AR-NW12A / Wavelength: 1 Å |
| Detector | Type: ADSC QUANTUM 210 / Detector: CCD / Date: May 24, 2005 |
| Radiation | Monochromator: Numerical link type Si(111) double crystal monochromator Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.6→50 Å / Num. obs: 11450 / % possible obs: 94.5 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 |
| Reflection shell | Resolution: 1.6→1.66 Å / % possible all: 64.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2DS5 Resolution: 1.6→30 Å / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→30 Å
| ||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||
| LS refinement shell | Resolution: 1.6→1.63 Å / Rfactor Rfree error: 0.0124
|
 Movie
Movie Controller
Controller













 PDBj
PDBj




