[English] 日本語
 Yorodumi
Yorodumi- PDB-2a0b: HISTIDINE-CONTAINING PHOSPHOTRANSFER DOMAIN OF ARCB FROM ESCHERIC... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2a0b | ||||||
|---|---|---|---|---|---|---|---|
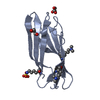
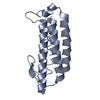
| Title | HISTIDINE-CONTAINING PHOSPHOTRANSFER DOMAIN OF ARCB FROM ESCHERICHIA COLI | ||||||
 Components Components | HPT DOMAIN | ||||||
 Keywords Keywords | SENSORY TRANSDUCTION / HISTIDINE KINASE / PHOSPHOTRANSFER / TWO-COMPONENT SYSTEM / FOUR-HELIX BUNDLE | ||||||
| Function / homology |  Function and homology information Function and homology informationpeptidyl-histidine phosphorylation / plasma membrane => GO:0005886 / response to oxygen levels / phosphorelay sensor kinase activity / histidine kinase / phosphoprotein phosphatase activity / protein autophosphorylation / regulation of DNA-templated transcription / signal transduction / ATP binding ...peptidyl-histidine phosphorylation / plasma membrane => GO:0005886 / response to oxygen levels / phosphorelay sensor kinase activity / histidine kinase / phosphoprotein phosphatase activity / protein autophosphorylation / regulation of DNA-templated transcription / signal transduction / ATP binding / plasma membrane / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.57 Å X-RAY DIFFRACTION / Resolution: 1.57 Å | ||||||
 Authors Authors | Kato, M. / Mizuno, T. / Shimizu, T. / Hakoshima, T. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 1999 Journal: Acta Crystallogr.,Sect.D / Year: 1999Title: Refined structure of the histidine-containing phosphotransfer (HPt) domain of the anaerobic sensor kinase ArcB from Escherichia coli at 1.57 A resolution. Authors: Kato, M. / Mizuno, T. / Shimizu, T. / Hakoshima, T. #1:  Journal: Cell(Cambridge,Mass.) / Year: 1997 Journal: Cell(Cambridge,Mass.) / Year: 1997Title: Insights Into Multistep Phosphorelay from the Crystal Structure of the C-Terminal Hpt Domain of Arcb Authors: Kato, M. / Mizuno, T. / Shimizu, T. / Hakoshima, T. #2:  Journal: Acta Crystallogr.,Sect.D / Year: 1996 Journal: Acta Crystallogr.,Sect.D / Year: 1996Title: Crystallization and Preliminary X-Ray Analysis of a Histidine Kinase Domain of the Anaerobic Sensor Protein Arcb from Escherichia Coli Authors: Kato, M. / Ishige, K. / Mizuno, T. / Shimizu, T. / Hakoshima, T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2a0b.cif.gz 2a0b.cif.gz | 40.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2a0b.ent.gz pdb2a0b.ent.gz | 27.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2a0b.json.gz 2a0b.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/a0/2a0b https://data.pdbj.org/pub/pdb/validation_reports/a0/2a0b ftp://data.pdbj.org/pub/pdb/validation_reports/a0/2a0b ftp://data.pdbj.org/pub/pdb/validation_reports/a0/2a0b | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14013.939 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   References: UniProt: P22763, UniProt: P0AEC3*PLUS, Transferases; Transferring phosphorus-containing groups; Phosphotransferases with a nitrogenous group as acceptor |
|---|---|
| #2: Chemical | ChemComp-ZN / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.1 Å3/Da / Density % sol: 41 % / Description: THE FURTHER REFINEMENT OF 1A0B | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 4.1 Details: PROTEIN WAS CRYSTALLIZED FROM 12.5% PEGMME 550, 5 MM ZNSO4, 50 MM ACETIC ACID/SODIUM ACETATE BUFFER, PH 4.1 | ||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 277 K / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 277 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 |
| Detector | Type: RIGAKU RAXIS IIC / Detector: IMAGE PLATE / Date: Sep 1, 1997 / Details: COLLIMATOR |
| Radiation | Monochromator: GRAPHITE(002) / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Highest resolution: 1.57 Å / Num. obs: 16338 / % possible obs: 94.7 % / Observed criterion σ(I): 1 / Redundancy: 6 % / Biso Wilson estimate: 26.8 Å2 / Rmerge(I) obs: 0.049 / Rsym value: 0.049 / Net I/σ(I): 66 |
| Reflection shell | Resolution: 1.57→1.7 Å / Redundancy: 5.1 % / Rmerge(I) obs: 0.19 / Rsym value: 0.19 / % possible all: 89 |
| Reflection | *PLUS % possible obs: 95 % / Num. measured all: 98202 |
| Reflection shell | *PLUS % possible obs: 89 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.57→6 Å / Cross valid method: THROUGHOUT / σ(F): 1 / Details: BULK SOLVENT MODEL USED
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 29.5 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.17 Å / Luzzati d res low obs: 6 Å / Luzzati sigma a obs: 0.17 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.57→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: REFMAC / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Num. reflection obs: 16036 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Highest resolution: 1.57 Å / Lowest resolution: 1.63 Å / Rfactor Rfree: 0.299 / Rfactor Rwork: 0.277 |
 Movie
Movie Controller
Controller










 PDBj
PDBj









