+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1z1z | ||||||
|---|---|---|---|---|---|---|---|
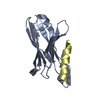
| Title | NMR structure of the gpu tail protein from lambda bacteriophage | ||||||
 Components Components | Minor tail protein U | ||||||
 Keywords Keywords | VIRAL PROTEIN / mixed alpha-beta / tail protein / bacteriophage | ||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont genome ejection through host cell envelope, long flexible tail mechanism / viral tail assembly / virus tail / host cell cytoplasm Similarity search - Function | ||||||
| Biological species |  Enterobacteria phage lambda (virus) Enterobacteria phage lambda (virus) | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
 Authors Authors | Edmonds, L. / Maxwell, K. / Davidson, A. / Donaldson, L.W. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2007 Journal: J.Mol.Biol. / Year: 2007Title: The NMR structure of the gpU tail-terminator protein from bacteriophage lambda: identification of sites contributing to Mg(II)-mediated oligomerization and biological function. Authors: Edmonds, L. / Liu, A. / Kwan, J.J. / Avanessy, A. / Caracoglia, M. / Yang, I. / Maxwell, K.L. / Rubenstein, J. / Davidson, A.R. / Donaldson, L.W. #1:  Journal: To be Published Journal: To be PublishedTitle: NMR assignment of the gpu tail protein from lambda bacteriophage Authors: Edmonds, L. / Thirumoorthy, R.T. / Liu, A. / Davidson, A. / Donaldson, L.W. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1z1z.cif.gz 1z1z.cif.gz | 50 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1z1z.ent.gz pdb1z1z.ent.gz | 36.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1z1z.json.gz 1z1z.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/z1/1z1z https://data.pdbj.org/pub/pdb/validation_reports/z1/1z1z ftp://data.pdbj.org/pub/pdb/validation_reports/z1/1z1z ftp://data.pdbj.org/pub/pdb/validation_reports/z1/1z1z | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 14659.124 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Enterobacteria phage lambda (virus) / Genus: Lambda-like viruses Enterobacteria phage lambda (virus) / Genus: Lambda-like virusesDescription: The full gpu gene was inserted into the NdeI and BamHI sites of pET15b resulting in an amino terminally hexahistidine tagged fusion protein with an intervening thrombin protease cleavage site Gene: gpu / Plasmid: pET15b / Species (production host): Escherichia coli / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||
| NMR details | Text: This structure was determined using standard 3D heteronuclear techniques |
- Sample preparation
Sample preparation
| Details | Contents: 0.9 mM gpU, U-15N,13C, 10 mM Tris-d11, pH 7.8, 50 mM potassium chloride, 0.02% sodium azide, 10% D2O, 90% H2O Solvent system: 10% D2O, 90% H2O |
|---|---|
| Sample conditions | Ionic strength: 50 mM KCl / pH: 7.8 / Pressure: 1 atm / Temperature: 293 K |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Radiation wavelength | Relative weight: 1 | |||||||||||||||
| NMR spectrometer |
|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 1 Details: 1180 NOE distance restraints (876 intramolecular, 505 short-range, 128 medium range, 371 long-range), 42 pairs of hydrogen bond distance restraints, 106 pairs of phi/psi dihedral angle ...Details: 1180 NOE distance restraints (876 intramolecular, 505 short-range, 128 medium range, 371 long-range), 42 pairs of hydrogen bond distance restraints, 106 pairs of phi/psi dihedral angle restraints and 41 amide residual dipolar coupling restraints were incorporated into the structure calculation. NOE calibration was performed with CANDID module of CYANA. Initially, 200 structures were calcuated with XPLOR-NIH. From this ensemble, 50 low-energy structure were refined in water using the XPLOR-NIH protocol by C. Spronk. The top 20 structures by energy had a backbone RMSD of 0.96 +/- 0.16 A on the secondary structure elements. In addition to residues 1-3 and 128-131 at the termini, a large loop spanning residues 45-60 is disordered. | ||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller













 PDBj
PDBj HSQC
HSQC