[English] 日本語
 Yorodumi
Yorodumi- PDB-1uqg: SELF-COMPLEMENTARY DNA 5'-D(CGCGCG)2, NMR, MINIMIZED AVERAGE STRUCTURE -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1uqg | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | SELF-COMPLEMENTARY DNA 5'-D(CGCGCG)2, NMR, MINIMIZED AVERAGE STRUCTURE | ||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords KeywordsDNA / DEOXYRIBONUCLEIC ACID | Function / homology | DNA |  Function and homology information Function and homology informationMethod | SOLUTION NMR |  Authors AuthorsLam, S.L. / Au-Yeung, S.C.F. |  Citation Citation Journal: J.Mol.Biol. / Year: 1997 Journal: J.Mol.Biol. / Year: 1997Title: Sequence-specific local structural variations in solution structures of d(CGXX'CG)2 and d(CAXX'TG)2 self-complementary deoxyribonucleic acids. Authors: Lam, S.L. / Au-Yeung, S.C. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1uqg.cif.gz 1uqg.cif.gz | 13.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1uqg.ent.gz pdb1uqg.ent.gz | 8.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1uqg.json.gz 1uqg.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/uq/1uqg https://data.pdbj.org/pub/pdb/validation_reports/uq/1uqg ftp://data.pdbj.org/pub/pdb/validation_reports/uq/1uqg ftp://data.pdbj.org/pub/pdb/validation_reports/uq/1uqg | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 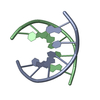 1bufC  1uqaC  1uqbC  1uqcC  1uqdC 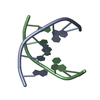 1uqeC 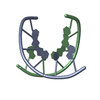 1uqfC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: DNA chain | Mass: 1810.205 Da / Num. of mol.: 2 / Source method: obtained synthetically Details: IN 10 MM SODIUM PHOSPHATE BUFFER AT 10 DEG. CELSIUS, 6.2 MM DOUBLE STRANDED CONCENTRATION |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|---|
| NMR details | Text: IN 10 MM SODIUM PHOSPHATE BUFFER AT 10 DEGREES CELSIUS, 6.2 MM DOUBLE STRANDED CONCENTRATION |
- Sample preparation
Sample preparation
| Sample conditions | Temperature: 283 K |
|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
- Processing
Processing
| Software | Name:  AMBER / Classification: refinement AMBER / Classification: refinement |
|---|---|
| NMR software | Name:  Amber / Version: 4 Amber / Version: 4 Developer: PEARLMAN,CASE,CALDWELL,SEIBEL,SINGH,WEINER, KOLLMAN Classification: refinement |
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller




 PDBj
PDBj

