+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 1sf0 | ||||||
|---|---|---|---|---|---|---|---|
| タイトル | BACKBONE SOLUTION STRUCTURE OF MIXED ALPHA/BETA PROTEIN PF1061 | ||||||
 要素 要素 | hypothetical protein PF1061 | ||||||
 キーワード キーワード | STRUCTURAL GENOMICS / UNKNOWN FUNCTION / RESIDUAL DIPOLAR COUPLINGS / PSI / Protein Structure Initiative / Southeast Collaboratory for Structural Genomics / SECSG | ||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報 | ||||||
| 生物種 |   Pyrococcus furiosus (古細菌) Pyrococcus furiosus (古細菌) | ||||||
| 手法 | 溶液NMR / RDC DIRECTED FRAGMENT ASSEMBLY | ||||||
 データ登録者 データ登録者 | Prestegard, J.H. / Mayer, K.L. / Valafar, H. / Southeast Collaboratory for Structural Genomics (SECSG) | ||||||
 引用 引用 |  ジャーナル: J.STRUCT.FUNCT.GENOM. / 年: 2005 ジャーナル: J.STRUCT.FUNCT.GENOM. / 年: 2005タイトル: Backbone solution structures of proteins using residual dipolar couplings: Application to a novel structural genomics target. 著者: Valafar, H. / Mayer, K.L. / Bougault, C.M. / Leblond, P.D. / Jenney, F.E. / Brereton, P.S. / Adams, M.W. / Prestegard, J.H. | ||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  1sf0.cif.gz 1sf0.cif.gz | 22.8 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb1sf0.ent.gz pdb1sf0.ent.gz | 11.8 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  1sf0.json.gz 1sf0.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| 文書・要旨 |  1sf0_validation.pdf.gz 1sf0_validation.pdf.gz | 319.1 KB | 表示 |  wwPDB検証レポート wwPDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  1sf0_full_validation.pdf.gz 1sf0_full_validation.pdf.gz | 321.4 KB | 表示 | |
| XML形式データ |  1sf0_validation.xml.gz 1sf0_validation.xml.gz | 2.7 KB | 表示 | |
| CIF形式データ |  1sf0_validation.cif.gz 1sf0_validation.cif.gz | 3.3 KB | 表示 | |
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/sf/1sf0 https://data.pdbj.org/pub/pdb/validation_reports/sf/1sf0 ftp://data.pdbj.org/pub/pdb/validation_reports/sf/1sf0 ftp://data.pdbj.org/pub/pdb/validation_reports/sf/1sf0 | HTTPS FTP |
-関連構造データ
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 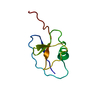
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR アンサンブル |
|
- 要素
要素
| #1: タンパク質 | 分子量: 8610.884 Da / 分子数: 1 / 由来タイプ: 組換発現 / 由来: (組換発現)   Pyrococcus furiosus (古細菌) / 株: DSM 3638 / 遺伝子: PF1061 / プラスミド: PET24D BAM / 発現宿主: Pyrococcus furiosus (古細菌) / 株: DSM 3638 / 遺伝子: PF1061 / プラスミド: PET24D BAM / 発現宿主:  |
|---|
-実験情報
-実験
| 実験 | 手法: 溶液NMR | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR実験 |
| ||||||||||||||||||||||||||||||||||||||||||||
| NMR実験の詳細 | Text: THIS STRUCTURE WAS DETERMINED USING PREDOMINANTLY RESIDUAL DIPOLAR COUPLINGS FROM BACKBONE ATOM PAIRS. IT IS A BACKBONE STRUCTURE MODELED AS AN ALA-GLY-PRO POLYPEPTIDE. |
- 試料調製
試料調製
| 詳細 |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 試料状態 |
|
-NMR測定
| 放射 | プロトコル: SINGLE WAVELENGTH / 単色(M)・ラウエ(L): M / 散乱光タイプ: x-ray | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 放射波長 | 相対比: 1 | |||||||||||||||
| NMRスペクトロメーター |
|
- 解析
解析
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 精密化 | 手法: RDC DIRECTED FRAGMENT ASSEMBLY / ソフトェア番号: 1 詳細: RDCS WERE USED IN THE INITIAL ASSEMBLY OF FOUR FRAGMENTS. RDCS FROM TWO MEDIA WERE USED TO SET RELATIVE ORIENTATIONS OF THE FRAGMENTS. TRANSLATIONAL RELATIONSHIPS OF FRAGMENTS WERE DICTATED ...詳細: RDCS WERE USED IN THE INITIAL ASSEMBLY OF FOUR FRAGMENTS. RDCS FROM TWO MEDIA WERE USED TO SET RELATIVE ORIENTATIONS OF THE FRAGMENTS. TRANSLATIONAL RELATIONSHIPS OF FRAGMENTS WERE DICTATED BY SEQUENCE CONNECTIVITIES AND LONG-RANGE NOES. THE ASSEMBLED STRUCTURE WAS MINIMIZED USING A MOLECULAR FORCE FIELD AND RDC ERROR FUNCTION. A TOTAL OF 486 RESTRAINTS WERE USED: 380 RESIDUAL DIPOLAR COUPLING RESTRAINTS, 85 NOE RESTRAINTS (OF WHICH 64 WERE SEQUENTIAL, 11 SHORT-RANGE AND 10 LONG-RANGE), AND 21 DIHEDRAL RESTRAINTS. ALL SIDECHAIN ATOMS BEYOND CB ARE MISSING. | ||||||||||||||||||||
| 代表構造 | 選択基準: fewest violations | ||||||||||||||||||||
| NMRアンサンブル | コンフォーマー選択の基準: structures with the least restraint violations 計算したコンフォーマーの数: 1 / 登録したコンフォーマーの数: 1 |
 ムービー
ムービー コントローラー
コントローラー










 PDBj
PDBj




 HNCA
HNCA