+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1sf0 | ||||||
|---|---|---|---|---|---|---|---|
| Title | BACKBONE SOLUTION STRUCTURE OF MIXED ALPHA/BETA PROTEIN PF1061 | ||||||
 Components Components | hypothetical protein PF1061 | ||||||
 Keywords Keywords | STRUCTURAL GENOMICS / UNKNOWN FUNCTION / RESIDUAL DIPOLAR COUPLINGS / PSI / Protein Structure Initiative / Southeast Collaboratory for Structural Genomics / SECSG | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |   Pyrococcus furiosus (archaea) Pyrococcus furiosus (archaea) | ||||||
| Method | SOLUTION NMR / RDC DIRECTED FRAGMENT ASSEMBLY | ||||||
 Authors Authors | Prestegard, J.H. / Mayer, K.L. / Valafar, H. / Southeast Collaboratory for Structural Genomics (SECSG) | ||||||
 Citation Citation |  Journal: J.STRUCT.FUNCT.GENOM. / Year: 2005 Journal: J.STRUCT.FUNCT.GENOM. / Year: 2005Title: Backbone solution structures of proteins using residual dipolar couplings: Application to a novel structural genomics target. Authors: Valafar, H. / Mayer, K.L. / Bougault, C.M. / Leblond, P.D. / Jenney, F.E. / Brereton, P.S. / Adams, M.W. / Prestegard, J.H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1sf0.cif.gz 1sf0.cif.gz | 22.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1sf0.ent.gz pdb1sf0.ent.gz | 11.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1sf0.json.gz 1sf0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1sf0_validation.pdf.gz 1sf0_validation.pdf.gz | 319.1 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1sf0_full_validation.pdf.gz 1sf0_full_validation.pdf.gz | 321.4 KB | Display | |
| Data in XML |  1sf0_validation.xml.gz 1sf0_validation.xml.gz | 2.7 KB | Display | |
| Data in CIF |  1sf0_validation.cif.gz 1sf0_validation.cif.gz | 3.3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/sf/1sf0 https://data.pdbj.org/pub/pdb/validation_reports/sf/1sf0 ftp://data.pdbj.org/pub/pdb/validation_reports/sf/1sf0 ftp://data.pdbj.org/pub/pdb/validation_reports/sf/1sf0 | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 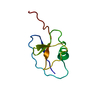
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 8610.884 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Pyrococcus furiosus (archaea) / Strain: DSM 3638 / Gene: PF1061 / Plasmid: PET24D BAM / Production host: Pyrococcus furiosus (archaea) / Strain: DSM 3638 / Gene: PF1061 / Plasmid: PET24D BAM / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||||||||||
| NMR details | Text: THIS STRUCTURE WAS DETERMINED USING PREDOMINANTLY RESIDUAL DIPOLAR COUPLINGS FROM BACKBONE ATOM PAIRS. IT IS A BACKBONE STRUCTURE MODELED AS AN ALA-GLY-PRO POLYPEPTIDE. |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions |
|
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Radiation wavelength | Relative weight: 1 | |||||||||||||||
| NMR spectrometer |
|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: RDC DIRECTED FRAGMENT ASSEMBLY / Software ordinal: 1 Details: RDCS WERE USED IN THE INITIAL ASSEMBLY OF FOUR FRAGMENTS. RDCS FROM TWO MEDIA WERE USED TO SET RELATIVE ORIENTATIONS OF THE FRAGMENTS. TRANSLATIONAL RELATIONSHIPS OF FRAGMENTS WERE DICTATED ...Details: RDCS WERE USED IN THE INITIAL ASSEMBLY OF FOUR FRAGMENTS. RDCS FROM TWO MEDIA WERE USED TO SET RELATIVE ORIENTATIONS OF THE FRAGMENTS. TRANSLATIONAL RELATIONSHIPS OF FRAGMENTS WERE DICTATED BY SEQUENCE CONNECTIVITIES AND LONG-RANGE NOES. THE ASSEMBLED STRUCTURE WAS MINIMIZED USING A MOLECULAR FORCE FIELD AND RDC ERROR FUNCTION. A TOTAL OF 486 RESTRAINTS WERE USED: 380 RESIDUAL DIPOLAR COUPLING RESTRAINTS, 85 NOE RESTRAINTS (OF WHICH 64 WERE SEQUENTIAL, 11 SHORT-RANGE AND 10 LONG-RANGE), AND 21 DIHEDRAL RESTRAINTS. ALL SIDECHAIN ATOMS BEYOND CB ARE MISSING. | ||||||||||||||||||||
| NMR representative | Selection criteria: fewest violations | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations Conformers calculated total number: 1 / Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller










 PDBj
PDBj




 HNCA
HNCA