[English] 日本語
 Yorodumi
Yorodumi- PDB-1qfo: N-TERMINAL DOMAIN OF SIALOADHESIN (MOUSE) IN COMPLEX WITH 3'SIALY... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qfo | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
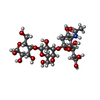
| Title | N-TERMINAL DOMAIN OF SIALOADHESIN (MOUSE) IN COMPLEX WITH 3'SIALYLLACTOSE | |||||||||
 Components Components | PROTEIN (SIALOADHESIN) | |||||||||
 Keywords Keywords | IMMUNE SYSTEM / IMMUNOGLOBULIN SUPERFAMILY / CARBOHYDRATE BINDING | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of T cell apoptotic process / virion binding / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / positive regulation of extrinsic apoptotic signaling pathway / negative regulation of type I interferon production / positive regulation of type I interferon production / late endosome / carbohydrate binding / clathrin-dependent endocytosis of virus by host cell / early endosome ...positive regulation of T cell apoptotic process / virion binding / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / positive regulation of extrinsic apoptotic signaling pathway / negative regulation of type I interferon production / positive regulation of type I interferon production / late endosome / carbohydrate binding / clathrin-dependent endocytosis of virus by host cell / early endosome / cell adhesion / extracellular region / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.85 Å MOLECULAR REPLACEMENT / Resolution: 1.85 Å | |||||||||
 Authors Authors | May, A.P. / Robinson, R.C. / Vinson, M. / Crocker, P.R. / Jones, E.Y. | |||||||||
 Citation Citation |  Journal: Mol.Cell / Year: 1998 Journal: Mol.Cell / Year: 1998Title: Crystal structure of the N-terminal domain of sialoadhesin in complex with 3' sialyllactose at 1.85 A resolution. Authors: May, A.P. / Robinson, R.C. / Vinson, M. / Crocker, P.R. / Jones, E.Y. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qfo.cif.gz 1qfo.cif.gz | 88.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qfo.ent.gz pdb1qfo.ent.gz | 66.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qfo.json.gz 1qfo.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qf/1qfo https://data.pdbj.org/pub/pdb/validation_reports/qf/1qfo ftp://data.pdbj.org/pub/pdb/validation_reports/qf/1qfo ftp://data.pdbj.org/pub/pdb/validation_reports/qf/1qfo | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||
| 2 | 
| ||||||||||||
| 3 | 
| ||||||||||||
| Unit cell |
| ||||||||||||
| Noncrystallographic symmetry (NCS) | NCS oper:
|
- Components
Components
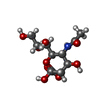
| #1: Protein | Mass: 13319.107 Da / Num. of mol.: 3 / Fragment: N-TERMINAL SIALIC ACID-BINDING DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Polysaccharide | #3: Sugar | ChemComp-SIA / | #4: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.54 Å3/Da / Density % sol: 52 % | |||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 5.6 Details: 30 % (W/V) PEG 4000, 10 MM DTT, 0.1M SODIUM HEPES PH 5.6, 5MG/ML PROTEIN, 12.5MM 3'SIALYLLACTOSE | |||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 17 ℃ / Method: vapor diffusion, sitting drop | |||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 288 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Photon Factory Photon Factory  / Beamline: BL-6A / Wavelength: 1 / Beamline: BL-6A / Wavelength: 1 |
| Detector | Type: FUJI / Detector: IMAGE PLATE |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.85→20 Å / Num. all: 33750 / Num. obs: 33750 / % possible obs: 94.6 % / Observed criterion σ(I): 0 / Biso Wilson estimate: 18.9 Å2 / Rmerge(I) obs: 0.071 |
| Reflection shell | Resolution: 1.85→1.97 Å / % possible all: 84.24 |
| Reflection shell | *PLUS % possible obs: 84.2 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.85→20 Å / Rfactor Rfree error: 0.005 / Data cutoff high rms absF: 3314732.04 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 MOLECULAR REPLACEMENT / Resolution: 1.85→20 Å / Rfactor Rfree error: 0.005 / Data cutoff high rms absF: 3314732.04 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 48.52 Å2 / ksol: 0.339 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 29.5 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.85→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.85→1.97 Å / Rfactor Rfree error: 0.016 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 0.5 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj