[English] 日本語
 Yorodumi
Yorodumi- PDB-1pup: CRYSTAL STRUCTURE OF A PEPTIDE NUCLEIC ACID (PNA) DUPLEX AT 1.7 A... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1pup | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
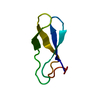
| Title | CRYSTAL STRUCTURE OF A PEPTIDE NUCLEIC ACID (PNA) DUPLEX AT 1.7 ANGSTROMS RESOLUTION | ||||||||||||||||||||||||
 Components Components | PNA (H-P(* Keywords KeywordsPEPTIDE NUCLEIC ACID / DNA ANALOGUE / DOUBLE HELIX | Function / homology | PEP_NUC |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SIR / Resolution: 1.7 Å SIR / Resolution: 1.7 Å  Authors AuthorsRasmussen, H. / Kastrup, J.S. |  Citation Citation Journal: Nat.Struct.Biol. / Year: 1997 Journal: Nat.Struct.Biol. / Year: 1997Title: Crystal structure of a peptide nucleic acid (PNA) duplex at 1.7 A resolution. Authors: Rasmussen, H. / Kastrup, J.S. / Nielsen, J.N. / Nielsen, J.M. / Nielsen, P.E. #1:  Journal: Science / Year: 1995 Journal: Science / Year: 1995Title: A Nucleic Acid Triple Helix Formed by a Peptide Nucleic Acid-DNA Complex Authors: Betts, L. / Josey, J.A. / Veal, J.M. / Jordan, S.R. #2:  Journal: Nat.Struct.Biol. / Year: 1996 Journal: Nat.Struct.Biol. / Year: 1996Title: Solution Structure of a Petide Nucleic Acid-DNA Duplex Authors: Eriksson, M. / Nielsen, P.E. #3:  Journal: Science / Year: 1994 Journal: Science / Year: 1994Title: NMR Solution Structure of a Peptide Nucleic Acid Complexed with RNA Authors: Brown, S.C. / Thomson, S.A. / Veal, J.M. / Davis, D.G. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1pup.cif.gz 1pup.cif.gz | 18.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1pup.ent.gz pdb1pup.ent.gz | 12.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1pup.json.gz 1pup.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pu/1pup https://data.pdbj.org/pub/pdb/validation_reports/pu/1pup ftp://data.pdbj.org/pub/pdb/validation_reports/pu/1pup ftp://data.pdbj.org/pub/pdb/validation_reports/pu/1pup | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Peptide nucleic acid | Mass: 1648.606 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source #2: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.51 Å3/Da / Density % sol: 51.01 % Description: DERIVATIVE DATA WERE COLLECTED AT EMBL, HAMBURG AT BEAMLINE X31 TO 1.9 A RESOLUTION. | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Method: vapor diffusion, hanging drop / pH: 7 / Details: pH 7.00, VAPOR DIFFUSION, HANGING DROP | |||||||||||||||||||||||||
| Components of the solutions |
| |||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃Details: drop solution was mixed with an equal volume of reservoir solution | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 285 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 ROTATING ANODE / Type: RIGAKU RU200 |
| Detector | Type: RIGAKU RAXIS II / Detector: IMAGE PLATE / Date: Sep 1, 1995 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
| Reflection | Resolution: 1.7→25 Å / Num. obs: 5222 / % possible obs: 77 % / Observed criterion σ(I): 1 / Redundancy: 1.9 % / Biso Wilson estimate: 15.8 Å2 / Rmerge(I) obs: 0.032 / Net I/σ(I): 13.1 |
| Reflection shell | Resolution: 1.7→1.9 Å / Redundancy: 1.9 % / Rmerge(I) obs: 0.058 / Mean I/σ(I) obs: 7.3 / % possible all: 37 |
| Reflection | *PLUS Highest resolution: 1.7 Å / Lowest resolution: 25 Å / % possible obs: 77 % / Redundancy: 1.9 % |
| Reflection shell | *PLUS Highest resolution: 1.7 Å / Lowest resolution: 1.9 Å / % possible obs: 37 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SIR / Resolution: 1.7→6 Å / Data cutoff high absF: 100000 / Data cutoff low absF: 0.1 / σ(F): 2 SIR / Resolution: 1.7→6 Å / Data cutoff high absF: 100000 / Data cutoff low absF: 0.1 / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 11.7 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine Biso |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.7→1.78 Å / Total num. of bins used: 8
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file | Serial no: 1 / Param file: PARAM.PNA / Topol file: TOPH_JULY96.PNA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.7 Å / Lowest resolution: 6 Å / σ(F): 2 / % reflection Rfree: 10 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor obs: 0.285 |
 Movie
Movie Controller
Controller









 PDBj
PDBj
