[English] 日本語
 Yorodumi
Yorodumi- PDB-1ppa: THE CRYSTAL STRUCTURE OF A LYSINE 49 PHOSPHOLIPASE A2 FROM THE VE... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ppa | ||||||
|---|---|---|---|---|---|---|---|
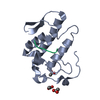
| Title | THE CRYSTAL STRUCTURE OF A LYSINE 49 PHOSPHOLIPASE A2 FROM THE VENOM OF THE COTTONMOUTH SNAKE AT 2.0 ANGSTROMS RESOLUTION | ||||||
 Components Components | PHOSPHOLIPASE A2 | ||||||
 Keywords Keywords | HYDROLASE(CARBOXYLIC ESTERASE) | ||||||
| Function / homology |  Function and homology information Function and homology information: / arachidonate secretion / lipid catabolic process / negative regulation of T cell proliferation / phospholipid metabolic process / phospholipid binding / toxin activity / calcium ion binding / extracellular region Similarity search - Function | ||||||
| Biological species |  Agkistrodon piscivorus piscivorus (Eastern cottonmouth) Agkistrodon piscivorus piscivorus (Eastern cottonmouth) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2 Å X-RAY DIFFRACTION / Resolution: 2 Å | ||||||
 Authors Authors | Holland, D.R. / Clancy, L.L. / Muchmore, S.W. / Rydel, T.J. / Einspahr, H.M. / Finzel, B.C. / Heinrikson, R.L. / Watenpaugh, K.D. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 1990 Journal: J.Biol.Chem. / Year: 1990Title: The crystal structure of a lysine 49 phospholipase A2 from the venom of the cottonmouth snake at 2.0-A resolution. Authors: Holland, D.R. / Clancy, L.L. / Muchmore, S.W. / Ryde, T.J. / Einspahr, H.M. / Finzel, B.C. / Heinrikson, R.L. / Watenpaugh, K.D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ppa.cif.gz 1ppa.cif.gz | 39.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ppa.ent.gz pdb1ppa.ent.gz | 27.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ppa.json.gz 1ppa.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pp/1ppa https://data.pdbj.org/pub/pdb/validation_reports/pp/1ppa ftp://data.pdbj.org/pub/pdb/validation_reports/pp/1ppa ftp://data.pdbj.org/pub/pdb/validation_reports/pp/1ppa | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Atom site foot note | 1: THE SIDE GROUP OF SER 74 IS DISORDERED SUCH THAT TWO DISCRETE 'OG' POSITIONS APPEAR IN THE ELECTRON DENSITY MAP. |
- Components
Components
| #1: Protein | Mass: 13987.321 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Agkistrodon piscivorus piscivorus (Eastern cottonmouth) Agkistrodon piscivorus piscivorus (Eastern cottonmouth)Species: Agkistrodon piscivorus / Strain: piscivorus / References: UniProt: P04361, phospholipase A2 |
|---|---|
| #2: Chemical | ChemComp-ANL / |
| #3: Water | ChemComp-HOH / |
| Has protein modification | Y |
| Nonpolymer details | THE SUBSTRATE CYLCLOHEXYLAMINE IS UNCERTAIN. IT WAS DESCRIBED AS A 'STRANGE DENSITY', MAYBE ...THE SUBSTRATE CYLCLOHEXY |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.63 Å3/Da / Density % sol: 53.28 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS pH: 9 / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | *PLUS Highest resolution: 2 Å / Num. obs: 10136 / Rmerge(I) obs: 0.08 |
- Processing
Processing
| Software | Name: CEDAR / Classification: refinement | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2→10 Å /
| ||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→10 Å
| ||||||||||||
| Software | *PLUS Name: CEDAR / Classification: refinement | ||||||||||||
| Refinement | *PLUS Highest resolution: 2 Å / Lowest resolution: 10 Å / Num. reflection obs: 10003 / Rfactor obs: 0.157 | ||||||||||||
| Solvent computation | *PLUS | ||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller











 PDBj
PDBj



