[English] 日本語
 Yorodumi
Yorodumi- PDB-1pfc: MOLECULAR-REPLACEMENT STRUCTURE OF GUINEA PIG IGG1 P*FC(PRIME) RE... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1pfc | ||||||
|---|---|---|---|---|---|---|---|
| Title | MOLECULAR-REPLACEMENT STRUCTURE OF GUINEA PIG IGG1 P*FC(PRIME) REFINED AT 3.1 ANGSTROMS RESOLUTION | ||||||
 Components Components | IGG1 PFC' FC | ||||||
 Keywords Keywords | IMMUNOGLOBULIN | ||||||
| Function / homology | Immunoglobulins / Immunoglobulin-like / Sandwich / Mainly Beta Function and homology information Function and homology information | ||||||
| Biological species |  Cavia porcellus (domestic guinea pig) Cavia porcellus (domestic guinea pig) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 3.125 Å X-RAY DIFFRACTION / Resolution: 3.125 Å | ||||||
 Authors Authors | Bryant, S.H. / Amzel, L.M. / Poljak, R.J. / Phizackerley, R.P. | ||||||
 Citation Citation | Journal: Acta Crystallogr.,Sect.B / Year: 1985 Title: Molecular-Replacement Structure of Guinea Pig Igg1 Pfc(Prime) Refined at 3.1 Angstroms Resolution Authors: Bryant, S.H. / Amzel, L.M. / Phizackerley, R.P. / Poljak, R.J. #1:  Journal: Thesis, Johns Hopkins University / Year: 1981 Journal: Thesis, Johns Hopkins University / Year: 1981Title: Crystallographic Refinement of Guinea Pig Iggi Pfc(Prime) Fragment at 3.125 Angstroms Authors: Bryant, S.H. #2:  Journal: Mol.Immunol. / Year: 1979 Journal: Mol.Immunol. / Year: 1979Title: Three Dimensional Structure of the P/Fc(Prime) Fragment of Guinea Pig Igg1 Authors: Phizackerley, R.P. / Wishner, B.C. / Bryant, S.H. / Amzel, L.M. / Lopez Decastro, J.A. / Poljak, R.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1pfc.cif.gz 1pfc.cif.gz | 36.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1pfc.ent.gz pdb1pfc.ent.gz | 21.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1pfc.json.gz 1pfc.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pf/1pfc https://data.pdbj.org/pub/pdb/validation_reports/pf/1pfc ftp://data.pdbj.org/pub/pdb/validation_reports/pf/1pfc ftp://data.pdbj.org/pub/pdb/validation_reports/pf/1pfc | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 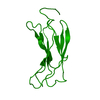
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 12690.335 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Cavia porcellus (domestic guinea pig) Cavia porcellus (domestic guinea pig) |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.46 Å3/Da / Density % sol: 50.1 % |
|---|---|
| Crystal grow | *PLUS Method: unknown |
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
- Processing
Processing
| Refinement | Resolution: 3.125→5 Å / Rfactor Rwork: 0.303 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement step | Cycle: LAST / Resolution: 3.125→5 Å
| ||||||||||||
| Refinement | *PLUS Rfactor obs: 0.303 | ||||||||||||
| Solvent computation | *PLUS | ||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller






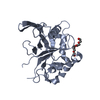





 PDBj
PDBj