[English] 日本語
 Yorodumi
Yorodumi- PDB-1pfb: Structural Basis for specific binding of polycomb chromodomain to... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1pfb | ||||||
|---|---|---|---|---|---|---|---|
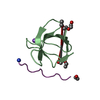
| Title | Structural Basis for specific binding of polycomb chromodomain to histone H3 methylated at K27 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | PEPTIDE BINDING PROTEIN / chromatin / histone methylation / polycomb / chromodomain | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of response to gamma radiation / specification of segmental identity, abdomen / RUNX1 interacts with co-factors whose precise effect on RUNX1 targets is not known / SUMOylation of DNA damage response and repair proteins / SUMOylation of RNA binding proteins / SUMOylation of transcription cofactors / Regulation of PTEN gene transcription / syncytial blastoderm mitotic cell cycle / polytene chromosome band / ventral cord development ...negative regulation of response to gamma radiation / specification of segmental identity, abdomen / RUNX1 interacts with co-factors whose precise effect on RUNX1 targets is not known / SUMOylation of DNA damage response and repair proteins / SUMOylation of RNA binding proteins / SUMOylation of transcription cofactors / Regulation of PTEN gene transcription / syncytial blastoderm mitotic cell cycle / polytene chromosome band / ventral cord development / PRC1 complex / Oxidative Stress Induced Senescence / polytene chromosome / anterior/posterior axis specification / histone H3K9me2/3 reader activity / neuron remodeling / neurogenesis / wound healing / structural constituent of chromatin / nucleosome / heterochromatin formation / sequence-specific DNA binding / protein heterodimerization activity / negative regulation of gene expression / negative regulation of DNA-templated transcription / chromatin binding / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / chromatin / nucleolus / negative regulation of transcription by RNA polymerase II / DNA binding / nucleus Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.4 Å MOLECULAR REPLACEMENT / Resolution: 1.4 Å | ||||||
 Authors Authors | Min, J.R. / Zhang, Y. / Xu, R.-M. | ||||||
 Citation Citation |  Journal: Genes Dev. / Year: 2003 Journal: Genes Dev. / Year: 2003Title: Structural basis for specific binding of Polycomb chromodomain to histone H3 methylated at Lys 27. Authors: Min, J.R. / Zhang, Y. / Xu, R.-M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1pfb.cif.gz 1pfb.cif.gz | 30.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1pfb.ent.gz pdb1pfb.ent.gz | 19.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1pfb.json.gz 1pfb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pf/1pfb https://data.pdbj.org/pub/pdb/validation_reports/pf/1pfb ftp://data.pdbj.org/pub/pdb/validation_reports/pf/1pfb ftp://data.pdbj.org/pub/pdb/validation_reports/pf/1pfb | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||||||||||||||||||||
| 2 | 
| |||||||||||||||||||||||||||
| Unit cell |
| |||||||||||||||||||||||||||
| Components on special symmetry positions |
|
- Components
Components
-Protein / Protein/peptide , 2 types, 2 molecules AB
| #1: Protein | Mass: 6806.826 Da / Num. of mol.: 1 / Fragment: chromodomain Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Protein/peptide | Mass: 1158.414 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: The peptide was chemically synthesized. It is naturally found in Strongylocentrotus purpuratus. References: UniProt: P06352 |
-Non-polymers , 4 types, 151 molecules 






| #3: Chemical | | #4: Chemical | ChemComp-BME / | #5: Chemical | ChemComp-ACY / | #6: Water | ChemComp-HOH / | |
|---|
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.03 Å3/Da / Density % sol: 59.39 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 290 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: 30% PEG8000, 200mM ammonium sulfate, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 290K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 16 ℃ / pH: 8 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X26C / Wavelength: 1.1 Å / Beamline: X26C / Wavelength: 1.1 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Nov 19, 2002 |
| Radiation | Monochromator: GRAPHITE / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.4→50 Å / Num. all: 19520 / Num. obs: 17050 / % possible obs: 86.8 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 |
| Reflection shell | Resolution: 1.4→1.45 Å / % possible all: 42.3 |
| Reflection | *PLUS Highest resolution: 1.4 Å / Num. measured all: 85538 / Rmerge(I) obs: 0.042 |
| Reflection shell | *PLUS Highest resolution: 1.4 Å / % possible obs: 42.3 % / Num. unique obs: 811 / Rmerge(I) obs: 0.195 / Mean I/σ(I) obs: 4.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.4→27.28 Å / σ(F): 0 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 1.4→27.28 Å / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.4→27.28 Å
| ||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.4 Å / Lowest resolution: 50 Å | ||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||
| LS refinement shell | *PLUS Highest resolution: 1.4 Å / Lowest resolution: 1.46 Å / Rfactor Rfree: 0.32 / Rfactor Rwork: 0.298 |
 Movie
Movie Controller
Controller








 PDBj
PDBj









