+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1npo | ||||||
|---|---|---|---|---|---|---|---|
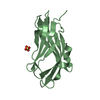
| Title | BOVINE NEUROPHYSIN II COMPLEX WITH OXYTOCIN | ||||||
 Components Components |
| ||||||
 Keywords Keywords | COMPLEX (HORMONE TRANSPORT/HORMONE) / COMPLEX (HORMONE TRANSPORT-HORMONE) / HYPOTHALAMUS / COMPLEX (HORMONE TRANSPORT-HORMONE) complex | ||||||
| Function / homology |  Function and homology information Function and homology informationmaternal aggressive behavior / neurohypophyseal hormone activity / V1A vasopressin receptor binding / vasoconstriction / neuropeptide hormone activity / positive regulation of systemic arterial blood pressure / positive regulation of vasoconstriction / secretory granule / intracellular signal transduction / extracellular space Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 3 Å X-RAY DIFFRACTION / Resolution: 3 Å | ||||||
 Authors Authors | Rose, J.P. / Wang, B.-C. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1996 Journal: Nat.Struct.Biol. / Year: 1996Title: Crystal structure of the neurophysin-oxytocin complex. Authors: Rose, J.P. / Wu, C.K. / Hsiao, C.D. / Breslow, E. / Wang, B.C. #1:  Journal: Proc.Natl.Acad.Sci.USA / Year: 1991 Journal: Proc.Natl.Acad.Sci.USA / Year: 1991Title: Crystal Structure of a Bovine Neurophysin II Dipeptide Complex at 2.8 A Determined from the Single-Wavelength Anomalous Scattering Signal of an Incorporated Iodine Atom Authors: Chen, L.Q. / Rose, J.P. / Breslow, E. / Yang, D. / Chang, W.R. / Furey Junior, W.F. / Sax, M. / Wang, B.C. #2:  Journal: Eur.J.Biochem. / Year: 1988 Journal: Eur.J.Biochem. / Year: 1988Title: Crystals of Modified Bovine Neurophysin II Authors: Rose, J.P. / Yang, D. / Yoo, C.S. / Sax, M. / Breslow, E. / Wang, B.C. #3:  Journal: J.Mol.Biol. / Year: 1979 Journal: J.Mol.Biol. / Year: 1979Title: Crystals of a Bovine Neurophysin II-Dipeptide Amide Complex Authors: Yoo, C.S. / Wang, B.C. / Sax, M. / Breslow, E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1npo.cif.gz 1npo.cif.gz | 43 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1npo.ent.gz pdb1npo.ent.gz | 31.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1npo.json.gz 1npo.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/np/1npo https://data.pdbj.org/pub/pdb/validation_reports/np/1npo ftp://data.pdbj.org/pub/pdb/validation_reports/np/1npo ftp://data.pdbj.org/pub/pdb/validation_reports/np/1npo | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 9890.252 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  | ||||
|---|---|---|---|---|---|
| #2: Protein/peptide | Mass: 1010.188 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source #3: Protein | | Mass: 9832.150 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.92 Å3/Da / Density % sol: 58 % / Description: DATA WAS COLLECTED IN 2 ORIENTATIONS. | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 6.8 / Details: pH 6.8 | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 18 K / pH: 6 / Method: batch method | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 295 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 |
| Detector | Type: SIEMENS-NICOLET X100 / Detector: AREA DETECTOR / Date: Jun 6, 1990 / Details: SUPPER MIRRORS |
| Radiation | Monochromator: NI FILTER / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Highest resolution: 3 Å / Num. obs: 5924 / % possible obs: 99.99 % / Redundancy: 6.4 % / Biso Wilson estimate: 29.17 Å2 / Rmerge(I) obs: 0.056 / Net I/σ(I): 10.6 |
| Reflection shell | Resolution: 3→3.18 Å / Redundancy: 5.6 % / Rmerge(I) obs: 0.253 / Mean I/σ(I) obs: 1.7 / % possible all: 100 |
| Reflection | *PLUS Num. all: 5928 / Num. measured all: 38324 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 3→8 Å / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 30.13 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 3→3.12 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj