+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 1mk6 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | SOLUTION STRUCTURE OF THE 8,9-DIHYDRO-8-(N7-GUANYL)-9-HYDROXY-AFLATOXIN B1 ADDUCT MISPAIRED WITH DEOXYADENOSINE | |||||||||
 要素 要素 |
| |||||||||
 キーワード キーワード | DNA / AFLATOXIN B1- Guanine Adduct Opposite an Adenine / mimicking GA Transition | |||||||||
| 機能・相同性 | 8,9-DIHYDRO-9-HYDROXY-AFLATOXIN B1 / DNA 機能・相同性情報 機能・相同性情報 | |||||||||
| 生物種 | synthetic construct (人工物) | |||||||||
| 手法 | 溶液NMR / distance geometry : MardiGras; simulated annealing, molecular dynamics : XPLOR; Average structure Calculation Addition of Solvent, and Counterions : AMBER; simulated annealing, Molecular Dynamics matrix relaxation : CORMA; | |||||||||
 データ登録者 データ登録者 | Giri, I. / Johnston, D.S. / Stone, M.P. | |||||||||
 引用 引用 |  ジャーナル: Biochemistry / 年: 2002 ジャーナル: Biochemistry / 年: 2002タイトル: MISPAIRING OF THE 8,9-DIHYDRO-8-(N7-GUANYL)-9-HYDROXY-AFLATOXIN B1 ADDUCT WITH DEOXYADENOSINE RESULTS IN EXTRUSION OF THE MISMATCHED DA TOWARD THE MAJOR GROOVE 著者: Giri, I. / Johnston, D.S. / Stone, M.P. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  1mk6.cif.gz 1mk6.cif.gz | 24.2 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb1mk6.ent.gz pdb1mk6.ent.gz | 15.6 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  1mk6.json.gz 1mk6.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/mk/1mk6 https://data.pdbj.org/pub/pdb/validation_reports/mk/1mk6 ftp://data.pdbj.org/pub/pdb/validation_reports/mk/1mk6 ftp://data.pdbj.org/pub/pdb/validation_reports/mk/1mk6 | HTTPS FTP |
|---|
-関連構造データ
| 関連構造データ | |
|---|---|
| 類似構造データ |
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 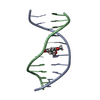
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR アンサンブル |
|
- 要素
要素
| #1: DNA鎖 | 分子量: 3003.993 Da / 分子数: 1 / 由来タイプ: 合成 / 由来: (合成) synthetic construct (人工物) |
|---|---|
| #2: DNA鎖 | 分子量: 3108.065 Da / 分子数: 1 / 由来タイプ: 合成 / 由来: (合成) synthetic construct (人工物) |
| #3: 化合物 | ChemComp-AFN / |
-実験情報
-実験
| 実験 | 手法: 溶液NMR | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR実験 |
| ||||||||||||||||||||||||
| NMR実験の詳細 | Text: 40 Structures were calculated starting from B and A form DNA. 20 closely convergent from both was averaged, and RMSD value was checked. The final averaged structure was calculated by averaging ...Text: 40 Structures were calculated starting from B and A form DNA. 20 closely convergent from both was averaged, and RMSD value was checked. The final averaged structure was calculated by averaging Final A and Final B, and after energy Minimization. This energy minimized structure was solvated, and explicit counterions were added. In all, 17 Na+ ions were added to neutralize the system using the Leap module in AMBER 6.0. The SHAKE algorithm constrained bonds involving protons to a tolerance of 0.0005 . A 1 fs time step was used. The rMD calculations were run for 1.4 ns, and coordinates were captured every 200 ps. The emergent structure from AMBER is Being reported |
- 試料調製
試料調製
| 詳細 | 内容: 80 OD of d(ACATCAFBGATCT)d(AGATAGATGT) solution in NMR Buffer 溶媒系: For observation of nonexchangeable protons, the sample was dissolved in 0.5 mL of 0.01 M sodium phosphate containing 0.1 M NaCl and 0.05 mM Na2EDTA at pD 7.4. The sample was dissolved in ...溶媒系: For observation of nonexchangeable protons, the sample was dissolved in 0.5 mL of 0.01 M sodium phosphate containing 0.1 M NaCl and 0.05 mM Na2EDTA at pD 7.4. The sample was dissolved in 99.96% D2O. For observation of exchangeable protons, the sample was dissolved in 9:1 H2O:D2O. Most experiments were performed at 5 C. Spectra of exchangeable protons were obtained at 0 C. | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 試料状態 |
| |||||||||||||||
| 結晶化 | *PLUS 手法: other / 詳細: NMR |
-NMR測定
| 放射 | プロトコル: SINGLE WAVELENGTH / 単色(M)・ラウエ(L): M | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 放射波長 | 相対比: 1 | ||||||||||||||||||||
| NMRスペクトロメーター |
|
- 解析
解析
| NMR software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 精密化 | 手法: distance geometry : MardiGras; simulated annealing, molecular dynamics : XPLOR; Average structure Calculation Addition of Solvent, and Counterions : AMBER; simulated annealing, Molecular ...手法: distance geometry : MardiGras; simulated annealing, molecular dynamics : XPLOR; Average structure Calculation Addition of Solvent, and Counterions : AMBER; simulated annealing, Molecular Dynamics matrix relaxation : CORMA; ソフトェア番号: 1 詳細: There were 329 experimental distance restraints derived from nonexchangeable 1H NOEs by MARDIGRAS. These consisted of 181 intranucleotide restraints, 110 internucleotide restraints, and 38 adduct-DNA restraints | ||||||||||||||||||||||||||||||||||||
| NMRアンサンブル | コンフォーマー選択の基準: Final Calculated Structure is Being Submitted. Back Calculated Structure is in Agreement with NOESY data. The calculation was performed in Presence of solvent ...コンフォーマー選択の基準: Final Calculated Structure is Being Submitted. Back Calculated Structure is in Agreement with NOESY data. The calculation was performed in Presence of solvent and counterions. Solvent, and Counterion co-ordinates are NOT being reported, only the Duplex DNA. Before Solvating and Addition of Counter IONS, 20 final structures were calculated using XPLOR. The final averaged energy minimized structure was solvated, and the counter Ions were added to it. Then MD was ran for 1.4 ns time scale to obtain final structure. 計算したコンフォーマーの数: 20 / 登録したコンフォーマーの数: 1 |
 ムービー
ムービー コントローラー
コントローラー







 PDBj
PDBj



 X-PLOR
X-PLOR