[English] 日本語
 Yorodumi
Yorodumi- PDB-1m9o: NMR structure of the first Zinc Binding domain of Nup475/TTP/TIS11 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1m9o | ||||||
|---|---|---|---|---|---|---|---|
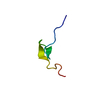
| Title | NMR structure of the first Zinc Binding domain of Nup475/TTP/TIS11 | ||||||
 Components Components | Tristetraproline | ||||||
 Keywords Keywords | METAL BINDING PROTEIN / Cys3His type zinc finger | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of keratinocyte apoptotic process / negative regulation of polynucleotide adenylyltransferase activity / nuclear-transcribed mRNA catabolic process, deadenylation-independent decay / Tristetraprolin (TTP, ZFP36) binds and destabilizes mRNA / protein-RNA sequence-specific adaptor activity / positive regulation of deadenylation-independent decapping of nuclear-transcribed mRNA / negative regulation of 3'-UTR-mediated mRNA stabilization / positive regulation of intracellular mRNA localization / negative regulation of erythrocyte differentiation / positive regulation of miRNA-mediated gene silencing ...regulation of keratinocyte apoptotic process / negative regulation of polynucleotide adenylyltransferase activity / nuclear-transcribed mRNA catabolic process, deadenylation-independent decay / Tristetraprolin (TTP, ZFP36) binds and destabilizes mRNA / protein-RNA sequence-specific adaptor activity / positive regulation of deadenylation-independent decapping of nuclear-transcribed mRNA / negative regulation of 3'-UTR-mediated mRNA stabilization / positive regulation of intracellular mRNA localization / negative regulation of erythrocyte differentiation / positive regulation of miRNA-mediated gene silencing / RNA destabilization / regulation of keratinocyte proliferation / regulation of keratinocyte differentiation / nuclear-transcribed mRNA poly(A) tail shortening / positive regulation of mRNA catabolic process / negative regulation of myeloid cell differentiation / 3'-UTR-mediated mRNA destabilization / negative regulation of viral transcription / miRNA-mediated gene silencing by inhibition of translation / regulation of tumor necrosis factor production / cellular response to granulocyte macrophage colony-stimulating factor stimulus / C-C chemokine binding / mRNA 3'-UTR AU-rich region binding / RNA polymerase binding / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / 3'-UTR-mediated mRNA stabilization / cellular response to glucocorticoid stimulus / negative regulation of interleukin-2 production / positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / response to starvation / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / p38MAPK cascade / mRNA catabolic process / mRNA transport / positive regulation of fat cell differentiation / heat shock protein binding / cellular response to fibroblast growth factor stimulus / 14-3-3 protein binding / regulation of mRNA stability / negative regulation of cytokine production involved in inflammatory response / cellular response to epidermal growth factor stimulus / mRNA 3'-UTR binding / P-body / response to wounding / negative regulation of inflammatory response / cellular response to tumor necrosis factor / cytoplasmic stress granule / MAPK cascade / cellular response to lipopolysaccharide / intracellular signal transduction / ribonucleoprotein complex / mRNA binding / regulation of transcription by RNA polymerase II / protein kinase binding / enzyme binding / negative regulation of transcription by RNA polymerase II / DNA binding / zinc ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR / CNS, dynamical annealing protocal with molecular dynamic scheme of torsion,torsion,cartesian. Initially only distance restraints from the three Cys to the zinc were restrained. Eventually the His was also restrained, angle restraints added to make the site tetrahedral. | ||||||
 Authors Authors | Amann, B.T. / Worthington, M.T. / Berg, J.M. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2003 Journal: Biochemistry / Year: 2003Title: A Cys3His Zinc-Binding Domain from Nup475/Tristetraproline: a Novel Fold with a Disklike Structure Authors: Amann, B.T. / Worthington, M.T. / Berg, J.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1m9o.cif.gz 1m9o.cif.gz | 293.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1m9o.ent.gz pdb1m9o.ent.gz | 237.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1m9o.json.gz 1m9o.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/m9/1m9o https://data.pdbj.org/pub/pdb/validation_reports/m9/1m9o ftp://data.pdbj.org/pub/pdb/validation_reports/m9/1m9o ftp://data.pdbj.org/pub/pdb/validation_reports/m9/1m9o | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 8972.193 Da / Num. of mol.: 1 / Fragment: zinc binding domain (Residues 91-163) / Mutation: Y143K Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Chemical | ChemComp-ZN / |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 1.7mM Nup475 with 2.2 Equivalent ZnCl2, 50mM d-Tris pH 5 Solvent system: 90% H20, 10% D20 and 100% D20 |
|---|---|
| Sample conditions | Ionic strength: no salt added / pH: 5.8 / Pressure: ambient / Temperature: 293 K |
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| NMR spectrometer | Type: Varian UNITYPLUS / Manufacturer: Varian / Model: UNITYPLUS / Field strength: 500 MHz |
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: CNS, dynamical annealing protocal with molecular dynamic scheme of torsion,torsion,cartesian. Initially only distance restraints from the three Cys to the zinc were restrained. Eventually the ...Method: CNS, dynamical annealing protocal with molecular dynamic scheme of torsion,torsion,cartesian. Initially only distance restraints from the three Cys to the zinc were restrained. Eventually the His was also restrained, angle restraints added to make the site tetrahedral. Software ordinal: 1 Details: 442 NOE restraints, 15 Torsional angle restraints. 15 zinc site restraints, 64 Carbon chemical shifts, and 174 proton chemical restraints. | ||||||||||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 30 / Conformers submitted total number: 23 |
 Movie
Movie Controller
Controller









 PDBj
PDBj


 HSQC
HSQC