[English] 日本語
 Yorodumi
Yorodumi- PDB-1lid: THE ADIPOCYTE LIPID-BINDING PROTEIN AT 1.6 ANGSTROMS RESOLUTION: ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1lid | ||||||
|---|---|---|---|---|---|---|---|
| Title | THE ADIPOCYTE LIPID-BINDING PROTEIN AT 1.6 ANGSTROMS RESOLUTION: CRYSTAL STRUCTURES OF THE APOPROTEIN AND WITH BOUND SATURATED AND UNSATURATED FATTY ACIDS | ||||||
 Components Components | ADIPOCYTE LIPID-BINDING PROTEIN | ||||||
 Keywords Keywords | LIPID BINDING PROTEIN / LIPID-BINDING PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationlipase activator activity / Triglyceride catabolism / lipid droplet formation / triglyceride catabolic process / hormone receptor binding / long-chain fatty acid transmembrane transporter activity / long-chain fatty acid binding / cellular response to lithium ion / white fat cell differentiation / long-chain fatty acid transport ...lipase activator activity / Triglyceride catabolism / lipid droplet formation / triglyceride catabolic process / hormone receptor binding / long-chain fatty acid transmembrane transporter activity / long-chain fatty acid binding / cellular response to lithium ion / white fat cell differentiation / long-chain fatty acid transport / fatty acid transport / lipid droplet / brown fat cell differentiation / cholesterol homeostasis / fatty acid binding / response to bacterium / fatty acid metabolic process / cellular response to tumor necrosis factor / positive regulation of inflammatory response / positive regulation of cold-induced thermogenesis / negative regulation of DNA-templated transcription / positive regulation of cell population proliferation / nucleoplasm / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.6 Å X-RAY DIFFRACTION / Resolution: 1.6 Å | ||||||
 Authors Authors | Zu, Z. / Bernlohr, D.A. / Banaszak, L.J. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 1993 Journal: J.Biol.Chem. / Year: 1993Title: The adipocyte lipid-binding protein at 1.6-A resolution. Crystal structures of the apoprotein and with bound saturated and unsaturated fatty acids. Authors: Xu, Z. / Bernlohr, D.A. / Banaszak, L.J. #1:  Journal: Biochemistry / Year: 1992 Journal: Biochemistry / Year: 1992Title: Crystal Structure of Recombinant Murine Adipocyte Lipid-Binding Protein Authors: Xu, Z. / Bernlohr, D.A. / Banaszak, L.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1lid.cif.gz 1lid.cif.gz | 39.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1lid.ent.gz pdb1lid.ent.gz | 27.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1lid.json.gz 1lid.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/li/1lid https://data.pdbj.org/pub/pdb/validation_reports/li/1lid ftp://data.pdbj.org/pub/pdb/validation_reports/li/1lid ftp://data.pdbj.org/pub/pdb/validation_reports/li/1lid | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 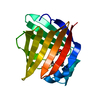
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14539.688 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Chemical | ChemComp-OLA / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.26 Å3/Da / Density % sol: 45.59 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS Temperature: 19 ℃ / pH: 6.9 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | *PLUS Highest resolution: 1.53 Å / Num. obs: 18975 / % possible obs: 73.3 % / Num. measured all: 82068 / Rmerge(I) obs: 0.0324 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.6→8 Å Details: SIDE CHAINS OF LYS 21, MET 35, LYS 37, AND LYS 79 ARE NOT RELIABLE.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.172 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: x_angle_d / Dev ideal: 2.5 |
 Movie
Movie Controller
Controller














 PDBj
PDBj








