+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1kf1 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure and Packing of Human Telomeric DNA | ||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / QUADRUPLEX DNA / DOUBLE CHAIN REVERSAL LOOP / HUMAN TELOMERE SEQUENCE / G(ANTI).G(ANTI).G(ANTI) / PARALLEL STRANDED | Function / homology | : / DNA / DNA (> 10) |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å MOLECULAR REPLACEMENT / Resolution: 2.1 Å  Authors AuthorsParkinson, G.N. / Lee, M.P.H. / Neidle, S. |  Citation Citation Journal: Nature / Year: 2002 Journal: Nature / Year: 2002Title: Crystal structure of parallel quadruplexes from human telomeric DNA. Authors: Parkinson, G.N. / Lee, M.P. / Neidle, S. #1:  Journal: Patent / Year: 2002 Journal: Patent / Year: 2002Title: Crystal Structure of G-Quadruplex Authors: Parkinson, G.N. / Lee, M.P.H. / Neidle, S. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1kf1.cif.gz 1kf1.cif.gz | 23.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1kf1.ent.gz pdb1kf1.ent.gz | 15.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1kf1.json.gz 1kf1.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kf/1kf1 https://data.pdbj.org/pub/pdb/validation_reports/kf/1kf1 ftp://data.pdbj.org/pub/pdb/validation_reports/kf/1kf1 ftp://data.pdbj.org/pub/pdb/validation_reports/kf/1kf1 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1k8pSC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 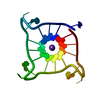
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: DNA chain | Mass: 6983.497 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: This sequence occurs naturally in humans | ||
|---|---|---|---|
| #2: Chemical | | #3: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.7 Å3/Da / Density % sol: 54.4 % | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 285 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: PEG 400, potassium iodide, potassium chloride, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 285K | ||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction | Mean temperature: 103 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-4 / Wavelength: 0.901 Å / Beamline: ID14-4 / Wavelength: 0.901 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Mar 31, 2001 |
| Radiation | Monochromator: Si 111 / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.901 Å / Relative weight: 1 |
| Reflection | Resolution: 2.1→20 Å / Num. all: 4495 / Num. obs: 4304 / % possible obs: 93.8 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 2 / Redundancy: 4 % / Rmerge(I) obs: 0.052 / Net I/σ(I): 50 |
| Reflection shell | Resolution: 2.1→2.17 Å / Redundancy: 4 % / Rmerge(I) obs: 0.117 / Mean I/σ(I) obs: 15 / Num. unique all: 440 / % possible all: 97.8 |
| Reflection | *PLUS Lowest resolution: 20 Å / Num. obs: 4595 / % possible obs: 98 % / Num. measured all: 50951 / Rmerge(I) obs: 0.052 |
| Reflection shell | *PLUS % possible obs: 97.8 % / Rmerge(I) obs: 0.117 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1K8P Resolution: 2.1→10 Å / Isotropic thermal model: isotropic / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 4 / Stereochemistry target values: parkinson et al.
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→10 Å
| ||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.09 Å / Rfactor Rwork: 0.296 | ||||||||||||||||||||
| Software | *PLUS Name: SHELXL / Version: 97 / Classification: refinement | ||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 10 Å / % reflection Rfree: 5 % / Rfactor all: 0.235 / Rfactor obs: 0.226 / Rfactor Rfree: 0.262 / Rfactor Rwork: 0.226 | ||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rwork: 0.296 |
 Movie
Movie Controller
Controller





 PDBj
PDBj









































