[English] 日本語
 Yorodumi
Yorodumi- PDB-1jia: STRUCTURE OF A BASIC PHOSPHOLIPASE A2 FROM AGKISTRODON HALYS PALL... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1jia | ||||||
|---|---|---|---|---|---|---|---|
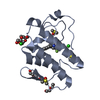
| Title | STRUCTURE OF A BASIC PHOSPHOLIPASE A2 FROM AGKISTRODON HALYS PALLAS AT 2.13A RESOLUTION | ||||||
 Components Components | PHOSPHOLIPASE A2 | ||||||
 Keywords Keywords | PHOSPHOLIPASE / PHOSPHOLIPASE A2 / AGKISTRODON HALYS PALLAS CRYSTAL STRUCTURE | ||||||
| Function / homology |  Function and homology information Function and homology informationphospholipase A2 / : / arachidonate secretion / lipid catabolic process / negative regulation of T cell proliferation / phospholipid metabolic process / phospholipid binding / toxin activity / calcium ion binding / extracellular region Similarity search - Function | ||||||
| Biological species |  Gloydius halys (Halys viper) Gloydius halys (Halys viper) | ||||||
| Method |  X-RAY DIFFRACTION / STRUCTURE REFINEMENT / Resolution: 2.13 Å X-RAY DIFFRACTION / STRUCTURE REFINEMENT / Resolution: 2.13 Å | ||||||
 Authors Authors | Zhao, K. / Lin, Z. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 1998 Journal: Acta Crystallogr.,Sect.D / Year: 1998Title: Structure of a basic phospholipase A2 from Agkistrodon halys Pallas at 2.13 A resolution. Authors: Zhao, K. / Song, S. / Lin, Z. / Zhou, Y. #1:  Journal: Shengwu Huaxue Zazhi / Year: 1997 Journal: Shengwu Huaxue Zazhi / Year: 1997Title: Preliminary Crystallographic Studies of a Basic Phospholipase A2 from the Venom of Agkistrodon Halys Pallas Authors: Zhao, K. / Song, S. / Lin, Z. / Zhou, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1jia.cif.gz 1jia.cif.gz | 65.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1jia.ent.gz pdb1jia.ent.gz | 47.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1jia.json.gz 1jia.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ji/1jia https://data.pdbj.org/pub/pdb/validation_reports/ji/1jia ftp://data.pdbj.org/pub/pdb/validation_reports/ji/1jia ftp://data.pdbj.org/pub/pdb/validation_reports/ji/1jia | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (0.10143, -0.99475, -0.01393), Vector: |
- Components
Components
| #1: Protein | Mass: 13923.168 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Source: (natural)  Gloydius halys (Halys viper) / Cellular location: EXTRACELLULAR / Secretion: VENOM / References: UniProt: O42187, phospholipase A2 Gloydius halys (Halys viper) / Cellular location: EXTRACELLULAR / Secretion: VENOM / References: UniProt: O42187, phospholipase A2#2: Chemical | #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.09 Å3/Da / Density % sol: 37 % Description: THE STRUCTURE OF BASIC PLA2 FROM AGKISTRODON HALYS PALLAS WAS DETERMINED BY MOLECULAR REPLACEMENT METHOD, USING R CHAIN OF PLA2 FROM THE VENOM OF WESTERN DIAMONDBACK RATTLESNAKE(2.5A) AS A SEARCH MODEL. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 9.5 Details: THE PROTEIN SOLUTIONS CONTAINED 10MM CA2+, 0.1M NACL, 5% PEG 4K IN 0.01M CHESS BUFFER (PH 9.5) AND AN ENZYME CONCENTRATION OF 11MG/ML; THE SOLUTION IN RESERVOIR CONTAINED 10% PEG 4K IN SAME ...Details: THE PROTEIN SOLUTIONS CONTAINED 10MM CA2+, 0.1M NACL, 5% PEG 4K IN 0.01M CHESS BUFFER (PH 9.5) AND AN ENZYME CONCENTRATION OF 11MG/ML; THE SOLUTION IN RESERVOIR CONTAINED 10% PEG 4K IN SAME BUFFER, ROOM TEMPERATURE OF 17 DEGREES C. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 280 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH2R / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RUH2R / Wavelength: 1.5418 |
| Detector | Type: SIEMENS-NICOLET X200B / Detector: AREA DETECTOR / Date: May 1, 1995 |
| Radiation | Monochromator: NICKEL / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.13→40 Å / Num. obs: 12001 / % possible obs: 86.1 % / Observed criterion σ(I): 0 / Redundancy: 2.5 % / Biso Wilson estimate: 26.24 Å2 / Rmerge(I) obs: 0.0459 / Net I/σ(I): 31.2 |
| Reflection shell | Resolution: 2.13→2.23 Å / Redundancy: 1.6 % / Rmerge(I) obs: 0.176 / Mean I/σ(I) obs: 4.9 / % possible all: 46.28 |
| Reflection | *PLUS Num. measured all: 29753 |
| Reflection shell | *PLUS % possible obs: 46.28 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: STRUCTURE REFINEMENT / Resolution: 2.13→6 Å Cross valid method: FULL REFINEMENT CONSISTING OF INITIAL MINIMIZATION, SLOW COOLING-SA 3000K, POSITIONAL AND B-FACTOR REFINEMENT. σ(F): 3 Details: DURING THE REFINEMENT, NO NCS RESTRAINTS WERE USED.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 25.41 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.25 Å / Luzzati d res low obs: 6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.13→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.13→2.22 Å / Total num. of bins used: 1
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor obs: 0.242 |
 Movie
Movie Controller
Controller











 PDBj
PDBj




