+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ip9 | ||||||
|---|---|---|---|---|---|---|---|
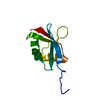
| Title | SOLUTION STRUCTURE OF THE PB1 DOMAIN OF BEM1P | ||||||
 Components Components | BEM1 PROTEIN | ||||||
 Keywords Keywords | SIGNALING PROTEIN / ubiquitin alpha/beta roll | ||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of vacuole fusion, non-autophagic / RHO GTPases Activate NADPH Oxidases / conjugation with cellular fusion / cell morphogenesis involved in conjugation with cellular fusion / site of polarized growth / PAR polarity complex / cellular bud site selection / incipient cellular bud site / maintenance of protein location / cellular bud tip ...positive regulation of vacuole fusion, non-autophagic / RHO GTPases Activate NADPH Oxidases / conjugation with cellular fusion / cell morphogenesis involved in conjugation with cellular fusion / site of polarized growth / PAR polarity complex / cellular bud site selection / incipient cellular bud site / maintenance of protein location / cellular bud tip / cellular bud neck / phosphatidylinositol-3-phosphate binding / regulation of Rho protein signal transduction / mating projection tip / cell cortex / molecular adaptor activity / cytoskeleton / protein-macromolecule adaptor activity / mitochondrion / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR / dynamical simulated annealing | ||||||
 Authors Authors | Terasawa, H. / Noda, Y. / Ito, T. / Hatanaka, H. / Ichikawa, S. / Ogura, K. / Sumimoto, H. / Inagaki, F. | ||||||
 Citation Citation |  Journal: EMBO J. / Year: 2001 Journal: EMBO J. / Year: 2001Title: Structure and ligand recognition of the PB1 domain: a novel protein module binding to the PC motif. Authors: Terasawa, H. / Noda, Y. / Ito, T. / Hatanaka, H. / Ichikawa, S. / Ogura, K. / Sumimoto, H. / Inagaki, F. #1:  Journal: Embo J. / Year: 2001 Journal: Embo J. / Year: 2001Title: Novel modular domain PB1 recognizes PC motif to mediate functional protein-protein interactions Authors: Ito, T. / Matsui, Y. / Ago, T. / Ota, K. / Sumimoto, H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ip9.cif.gz 1ip9.cif.gz | 586.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ip9.ent.gz pdb1ip9.ent.gz | 502.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ip9.json.gz 1ip9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ip/1ip9 https://data.pdbj.org/pub/pdb/validation_reports/ip/1ip9 ftp://data.pdbj.org/pub/pdb/validation_reports/ip/1ip9 ftp://data.pdbj.org/pub/pdb/validation_reports/ip/1ip9 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 9519.925 Da / Num. of mol.: 1 / Fragment: PB1 DOMAIN(RESIDUES 472-551) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Plasmid: PPROEX-HTA / Species (production host): Escherichia coli / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||
| NMR details | Text: The structure was determined using triple-resonance NMR spectroscopy. |
- Sample preparation
Sample preparation
| Details | Contents: 2mM Bem PB1 U-15N, 13C; 50mM potassium phosphate pH 6.3; 150mM sodium chloride; 1mM sodium azide; 90% H2O, 10% D2O Solvent system: 90% H2O, 10% D2O, 100% D2O |
|---|---|
| Sample conditions | Ionic strength: 50mM potassium phosphate, 150mM sodium chloride pH: 6.3 / Pressure: ambient / Temperature: 298 K |
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Radiation wavelength | Relative weight: 1 | |||||||||||||||
| NMR spectrometer |
|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: dynamical simulated annealing / Software ordinal: 1 Details: The structures are based on a total of 1334 restraints, 1269 are NOE-derived distance constraints, 65 dihedral angle restraints. | ||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 200 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller









 PDBj
PDBj

 X-PLOR
X-PLOR