+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1hmh | ||||||
|---|---|---|---|---|---|---|---|
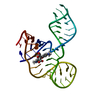
| Title | THREE-DIMENSIONAL STRUCTURE OF A HAMMERHEAD RIBOZYME | ||||||
 Components Components |
| ||||||
 Keywords Keywords | RIBOZYME / DNA-RNA HAMMERHEAD RIBOZYME / LOOP | ||||||
| Function / homology | DNA / DNA (> 10) / RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 2.6 Å SYNCHROTRON / Resolution: 2.6 Å | ||||||
 Authors Authors | Pley, H.W. / Flaherty, K.M. / McKay, D.B. | ||||||
 Citation Citation |  Journal: Nature / Year: 1994 Journal: Nature / Year: 1994Title: Three-dimensional structure of a hammerhead ribozyme. Authors: Pley, H.W. / Flaherty, K.M. / McKay, D.B. #1:  Journal: Nature / Year: 1994 Journal: Nature / Year: 1994Title: Model for an RNA Tertiary Interaction from the Structure of an Intermolecular Complex Between a GAAA Tetraloop and an RNA Helix Authors: Pley, H.W. / Flaherty, K.M. / McKay, D.B. #2:  Journal: Cell(Cambridge,Mass.) / Year: 1995 Journal: Cell(Cambridge,Mass.) / Year: 1995Title: The Crystal Structure of an All-RNA Hammerhead Ribozyme: a Proposed Mechanism for RNA Catalytic Cleavage Authors: Scott, W.G. / Finch, J.T. / Klug, A. #3:  Journal: J.Biol.Chem. / Year: 1993 Journal: J.Biol.Chem. / Year: 1993Title: Crystals of a Hammerhead Ribozyme Authors: Pley, H.W. / Lindes, D.S. / Deluca-Flaherty, C. / McKay, D.B. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1hmh.cif.gz 1hmh.cif.gz | 83.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1hmh.ent.gz pdb1hmh.ent.gz | 62.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1hmh.json.gz 1hmh.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/hm/1hmh https://data.pdbj.org/pub/pdb/validation_reports/hm/1hmh ftp://data.pdbj.org/pub/pdb/validation_reports/hm/1hmh ftp://data.pdbj.org/pub/pdb/validation_reports/hm/1hmh | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 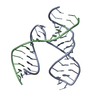
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 11030.680 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source #2: DNA chain | Mass: 3968.573 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 4.8 Å3/Da / Density % sol: 74.37 % | |||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 6 Details: pH 6.00, VAPOR DIFFUSION, HANGING DROP, temperature 277.00K | |||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| |||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 6 | |||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL7-1 / Beamline: BL7-1 |
|---|---|
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE |
| Radiation | Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
| Reflection | Highest resolution: 2.6 Å / Num. all: 135854 / Num. obs: 26816 |
- Processing
Processing
| Software | Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2.6→8 Å / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.6→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.6 Å / Lowest resolution: 8 Å / σ(F): 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller








 PDBj
PDBj































































