[English] 日本語
 Yorodumi
Yorodumi- PDB-1h5p: Solution structure of the human Sp100b SAND domain by heteronucle... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1h5p | ||||||
|---|---|---|---|---|---|---|---|
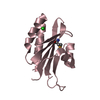
| Title | Solution structure of the human Sp100b SAND domain by heteronuclear NMR. | ||||||
 Components Components | NUCLEAR AUTOANTIGEN SP100-B | ||||||
 Keywords Keywords | NUCLEAR PROTEIN / TRANSCRIPTION / DNA BINDING / SP100B / SAND DOMAIN / KDWK / ANTIGEN / ALTERNATIVE SPLICING | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of Fas signaling pathway / regulation of viral process / maintenance of protein location / chromo shadow domain binding / response to type I interferon / negative regulation of protein export from nucleus / negative regulation of viral transcription / regulation of extrinsic apoptotic signaling pathway via death domain receptors / negative regulation of endothelial cell migration / type I interferon-mediated signaling pathway ...regulation of Fas signaling pathway / regulation of viral process / maintenance of protein location / chromo shadow domain binding / response to type I interferon / negative regulation of protein export from nucleus / negative regulation of viral transcription / regulation of extrinsic apoptotic signaling pathway via death domain receptors / negative regulation of endothelial cell migration / type I interferon-mediated signaling pathway / regulation of angiogenesis / response to retinoic acid / type II interferon-mediated signaling pathway / response to type II interferon / SUMOylation of DNA damage response and repair proteins / retinoic acid receptor signaling pathway / response to cytokine / nuclear periphery / telomere maintenance / DNA damage response, signal transduction by p53 class mediator / PML body / kinase binding / Interferon gamma signaling / transcription corepressor activity / DNA-binding transcription activator activity, RNA polymerase II-specific / RNA polymerase II-specific DNA-binding transcription factor binding / DNA-binding transcription factor activity, RNA polymerase II-specific / chromosome, telomeric region / protein dimerization activity / nuclear body / protein domain specific binding / negative regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / nucleolus / negative regulation of transcription by RNA polymerase II / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / DNA binding / nucleoplasm / identical protein binding / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species |  HOMO SAPIENS (human) HOMO SAPIENS (human) | ||||||
| Method | SOLUTION NMR / ARIA AMBIGUOUS DISTANCE RESTRAINTS SIMULATED ANNEALING WITH TORSION ANGLE DYNAMICS | ||||||
 Authors Authors | Bottomley, M.J. / Liu, Z. / Collard, M.W. / Huggenvik, J.I. / Gibson, T.J. / Sattler, M. | ||||||
 Citation Citation |  Journal: Nat. Struct. Biol. / Year: 2001 Journal: Nat. Struct. Biol. / Year: 2001Title: The SAND domain structure defines a novel DNA-binding fold in transcriptional regulation. Authors: Bottomley, M.J. / Collard, M.W. / Huggenvik, J.I. / Liu, Z. / Gibson, T.J. / Sattler, M. #1: Journal: J.Biol.Chem. / Year: 1999 Title: Nuclear Deaf-1-Related (Nudr) Protein Contains a Novel DNA Binding Domain and Represses Transcription of the Heterogeneous Nuclear Ribonucleoprotein A2/B1 Promoter Authors: Michelson, R.J. / Collard, M.W. / Ziemba, A.J. / Persinger, J. / Bartholomew, B. / Huggenvik, J.I. #2: Journal: Trends Biochem.Sci. / Year: 1998 Title: The Apeced Polyglandular Autoimmune Syndrome Protein, Aire-1, Contains the Sand Domain and is Probably a Transcription Factor Authors: Gibson, T.J. / Ramu, C. / Gemuend, C. / Aasland, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1h5p.cif.gz 1h5p.cif.gz | 344 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1h5p.ent.gz pdb1h5p.ent.gz | 289.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1h5p.json.gz 1h5p.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/h5/1h5p https://data.pdbj.org/pub/pdb/validation_reports/h5/1h5p ftp://data.pdbj.org/pub/pdb/validation_reports/h5/1h5p ftp://data.pdbj.org/pub/pdb/validation_reports/h5/1h5p | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11052.805 Da / Num. of mol.: 1 / Fragment: RESIDUES 595-688 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Plasmid: MODIFIED PET-24D HOMO SAPIENS (human) / Plasmid: MODIFIED PET-24DGene (production host): EXPRESSED AS FUSION PROTEIN WITH N-TERMINAL 6-HISTIDINE TAG AND A TEV PROTEASE CLEAVAGE SITE Production host:  |
|---|---|
| Sequence details | THE STRUCTURE IS FOR THE SPLICE VARIANT SP100B SAND DOMAIN. THE RESIDUES 685 -688 (RILE IN ISOFORM ...THE STRUCTURE IS FOR THE SPLICE VARIANT SP100B SAND DOMAIN. THE RESIDUES 685 -688 (RILE IN ISOFORM SP100-A) ARE (VTIK IN ISOFORM SP100-B) DIFFERENT IN THE TWO SPLICE VARIANTS MET A 594 IS A CLONING ARTIFACT. |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 1.2 MM PROTEIN, 20 MM SODIUM PHOSPHATE PH 6.5, 50 MM NACL, 4 MM DTT |
|---|---|
| Sample conditions | Ionic strength: 20 MM SODIUM PHOSPHATE, 50 MM SODIUM CHLORIDE pH: 6.5 / Pressure: 1 atm / Temperature: 295.0 K |
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: ARIA AMBIGUOUS DISTANCE RESTRAINTS SIMULATED ANNEALING WITH TORSION ANGLE DYNAMICS Software ordinal: 1 Details: DURING THE STRUCTURE CALCULATIONS, THE PROGRAM ARIA 1.0 (M.NILGES & S.I.O'DONOGHUE) WAS USED TO EMPLOY DISTANCE RESTRAINTS DERIVED FROM AMBIGUOUSLY ASSIGNED NOES. FROM A TOTAL OF 2120 NOES, ...Details: DURING THE STRUCTURE CALCULATIONS, THE PROGRAM ARIA 1.0 (M.NILGES & S.I.O'DONOGHUE) WAS USED TO EMPLOY DISTANCE RESTRAINTS DERIVED FROM AMBIGUOUSLY ASSIGNED NOES. FROM A TOTAL OF 2120 NOES, ABOUT 80% WERE MANUALLY ASSIGNED, THE REMAINDER WAS ASSIGNED BY ARIA. | ||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: ACCEPTABLE COVALENT GEOMETRY, FAVOURABLE NON-BOND ENERGY, FEWEST RESTRAINT VIOLATIONS, LOWEST OVERALL ENERGIES Conformers calculated total number: 100 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller











 PDBj
PDBj



 NMRPipe
NMRPipe