[English] 日本語
 Yorodumi
Yorodumi- PDB-1ey3: STRUCTURE OF ENOYL-COA HYDRATASE COMPLEXED WITH THE SUBSTRATE DAC-COA -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ey3 | ||||||
|---|---|---|---|---|---|---|---|
| Title | STRUCTURE OF ENOYL-COA HYDRATASE COMPLEXED WITH THE SUBSTRATE DAC-COA | ||||||
 Components Components | ENOYL-COA HYDRATASE | ||||||
 Keywords Keywords | LYASE / BETA-OXIDATION / CROTONASE / ENOYL-COA HYDRATASE / FATTY ACID METABOLISM / BETA-ELIMINATION / SYN-ADDITION / CONCERTED REACTION | ||||||
| Function / homology |  Function and homology information Function and homology informationBeta oxidation of lauroyl-CoA to decanoyl-CoA-CoA / Beta oxidation of decanoyl-CoA to octanoyl-CoA-CoA / Beta oxidation of octanoyl-CoA to hexanoyl-CoA / Beta oxidation of hexanoyl-CoA to butanoyl-CoA / Beta oxidation of butanoyl-CoA to acetyl-CoA / 3-hydroxypropionyl-CoA dehydratase activity / (2E)-butenoyl-CoA hydratase activity / Branched-chain amino acid catabolism / Delta3-Delta2-enoyl-CoA isomerase / delta(3)-delta(2)-enoyl-CoA isomerase activity ...Beta oxidation of lauroyl-CoA to decanoyl-CoA-CoA / Beta oxidation of decanoyl-CoA to octanoyl-CoA-CoA / Beta oxidation of octanoyl-CoA to hexanoyl-CoA / Beta oxidation of hexanoyl-CoA to butanoyl-CoA / Beta oxidation of butanoyl-CoA to acetyl-CoA / 3-hydroxypropionyl-CoA dehydratase activity / (2E)-butenoyl-CoA hydratase activity / Branched-chain amino acid catabolism / Delta3-Delta2-enoyl-CoA isomerase / delta(3)-delta(2)-enoyl-CoA isomerase activity / enoyl-CoA hydratase / enoyl-CoA hydratase activity / fatty acid beta-oxidation / mitochondrial matrix / mitochondrion Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.3 Å X-RAY DIFFRACTION / Resolution: 2.3 Å | ||||||
 Authors Authors | Bahnson, B.J. / Anderson, V.E. / Petsko, G.A. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2002 Journal: Biochemistry / Year: 2002Title: Structural mechanism of enoyl-CoA hydratase: three atoms from a single water are added in either an E1cb stepwise or concerted fashion. Authors: Bahnson, B.J. / Anderson, V.E. / Petsko, G.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ey3.cif.gz 1ey3.cif.gz | 312.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ey3.ent.gz pdb1ey3.ent.gz | 257.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ey3.json.gz 1ey3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1ey3_validation.pdf.gz 1ey3_validation.pdf.gz | 1.5 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1ey3_full_validation.pdf.gz 1ey3_full_validation.pdf.gz | 1.8 MB | Display | |
| Data in XML |  1ey3_validation.xml.gz 1ey3_validation.xml.gz | 62.7 KB | Display | |
| Data in CIF |  1ey3_validation.cif.gz 1ey3_validation.cif.gz | 84.6 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ey/1ey3 https://data.pdbj.org/pub/pdb/validation_reports/ey/1ey3 ftp://data.pdbj.org/pub/pdb/validation_reports/ey/1ey3 ftp://data.pdbj.org/pub/pdb/validation_reports/ey/1ey3 | HTTPS FTP |
-Related structure data
| Related structure data |  1dubS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Details | 6-subunits (A-F) come together as a dimer of trimers, forming a homo-hexamer, which is the biologically active form. |
- Components
Components
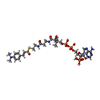
| #1: Protein | Mass: 28079.285 Da / Num. of mol.: 6 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Chemical | ChemComp-DAK / #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.41 Å3/Da / Density % sol: 48.76 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 7.3 Details: 8% PEG 4000, 100 mM sodium acetate, 75 mM sodium phosphate, 100 mM NaCl, 3 mM sodium azide, 3.5 mM DAC-CoA, pH 7.3, VAPOR DIFFUSION, HANGING DROP, temperature 298.0K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 25 ℃ | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 275 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 |
| Detector | Type: RIGAKU RAXIS II / Detector: IMAGE PLATE / Date: Aug 21, 1996 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→35 Å / Num. all: 65075 / Num. obs: 65075 / % possible obs: 94.8 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 4.9 % / Biso Wilson estimate: 40.8 Å2 / Rmerge(I) obs: 0.062 / Net I/σ(I): 11.9 |
| Reflection shell | Resolution: 2.3→2.38 Å / Redundancy: 1.64 % / Rmerge(I) obs: 0.413 / Num. unique all: 5554 / % possible all: 77.1 |
| Reflection | *PLUS Lowest resolution: 30 Å / % possible obs: 95 % |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Starting model: 1dub Resolution: 2.3→35 Å / σ(F): 0 / σ(I): 0 / Stereochemistry target values: protein_rep.param / Details: maximum likelihood target using amplitudes
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→35 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Classification: refinement | |||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.3 Å / Lowest resolution: 30 Å / σ(F): 0 / % reflection Rfree: 10 % / Rfactor obs: 0.18 | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller










 PDBj
PDBj