+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1edm | ||||||
|---|---|---|---|---|---|---|---|
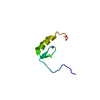
| Title | EPIDERMAL GROWTH FACTOR-LIKE DOMAIN FROM HUMAN FACTOR IX | ||||||
 Components Components | FACTOR IX | ||||||
 Keywords Keywords | COAGULATION FACTOR / EPIDERMAL GROWTH FACTOR / EGF / CALCIUM-BINDING / EGF-LIKE DOMAIN / STRUCTURE AND FUNCTION / HUMAN FACTOR IX | ||||||
| Function / homology |  Function and homology information Function and homology informationDefective F9 secretion / coagulation factor IXa / Defective gamma-carboxylation of F9 / Defective F9 activation / Defective factor IX causes thrombophilia / Defective cofactor function of FVIIIa variant / Defective F9 variant does not activate FX / : / zymogen activation / Protein hydroxylation ...Defective F9 secretion / coagulation factor IXa / Defective gamma-carboxylation of F9 / Defective F9 activation / Defective factor IX causes thrombophilia / Defective cofactor function of FVIIIa variant / Defective F9 variant does not activate FX / : / zymogen activation / Protein hydroxylation / Transport of gamma-carboxylated protein precursors from the endoplasmic reticulum to the Golgi apparatus / Gamma-carboxylation of protein precursors / Removal of aminoterminal propeptides from gamma-carboxylated proteins / : / Golgi lumen / blood coagulation / extracellular matrix / endopeptidase activity / endoplasmic reticulum lumen / serine-type endopeptidase activity / calcium ion binding / proteolysis / : / extracellular exosome / extracellular region / metal ion binding / plasma membrane Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MIR / Resolution: 1.5 Å MIR / Resolution: 1.5 Å | ||||||
 Authors Authors | Rao, Z. / Handford, P. / Mayhew, M. / Knott, V. / Brownlee, G.G. / Stuart, D. | ||||||
 Citation Citation |  Journal: Cell(Cambridge,Mass.) / Year: 1995 Journal: Cell(Cambridge,Mass.) / Year: 1995Title: The structure of a Ca(2+)-binding epidermal growth factor-like domain: its role in protein-protein interactions. Authors: Rao, Z. / Handford, P. / Mayhew, M. / Knott, V. / Brownlee, G.G. / Stuart, D. #1:  Journal: Acta Crystallogr.,Sect.D / Year: 1995 Journal: Acta Crystallogr.,Sect.D / Year: 1995Title: Crystallization of a Calcium-Binding Egf-Like Domain Authors: Rao, Z. / Handford, P. / Mayhew, M. / Knott, V. / Brownlee, G.G. / Stuart, D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1edm.cif.gz 1edm.cif.gz | 31.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1edm.ent.gz pdb1edm.ent.gz | 21.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1edm.json.gz 1edm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ed/1edm https://data.pdbj.org/pub/pdb/validation_reports/ed/1edm ftp://data.pdbj.org/pub/pdb/validation_reports/ed/1edm ftp://data.pdbj.org/pub/pdb/validation_reports/ed/1edm | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein/peptide | Mass: 4279.635 Da / Num. of mol.: 2 / Fragment: EPIDERMAL GROWTH FACTOR-LIKE DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Strain: RZ1032, XL1-BLUE, MC1061 / Gene: HUMAN FACTOR IX / Plasmid: PMA91 / Gene (production host): HUMAN FACTOR IX / Production host: Homo sapiens (human) / Strain: RZ1032, XL1-BLUE, MC1061 / Gene: HUMAN FACTOR IX / Plasmid: PMA91 / Gene (production host): HUMAN FACTOR IX / Production host:  #2: Chemical | #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.33 Å3/Da / Density % sol: 30 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 7.3 / Details: pH 7.3 | |||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 289 K / Method: vapor diffusion, sitting drop | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CHESS CHESS  / Beamline: F1 / Wavelength: 1.54 / Beamline: F1 / Wavelength: 1.54 |
| Detector | Type: SIEMENS / Detector: IMAGE PLATE / Date: Jan 1, 1992 |
| Radiation | Monochromator: Y / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.54 Å / Relative weight: 1 |
| Reflection | Resolution: 3.5→30 Å / Num. obs: 1196 / % possible obs: 97.6 % / Observed criterion σ(I): 0 / Redundancy: 3.5 % / Rmerge(I) obs: 0.058 |
| Reflection shell | Resolution: 1.5→1.65 Å / % possible all: 92 |
| Reflection | *PLUS Highest resolution: 1.5 Å / Num. obs: 13041 / % possible obs: 94.5 % / Redundancy: 3.7 % / Num. measured all: 48451 / Rmerge(I) obs: 0.102 |
| Reflection shell | *PLUS % possible obs: 92 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIR MIRStarting model: NMR MODEL Resolution: 1.5→30 Å / σ(F): 0 /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 17.5 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.5→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller










 PDBj
PDBj








