+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1dwu | ||||||
|---|---|---|---|---|---|---|---|
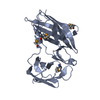
| Title | Ribosomal protein L1 | ||||||
 Components Components | RIBOSOMAL PROTEIN L1 | ||||||
 Keywords Keywords | RIBOSOMAL PROTEIN / RNA BINDING / PROTEIN SYNTHESIS | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of translation / large ribosomal subunit / tRNA binding / rRNA binding / structural constituent of ribosome / translation Similarity search - Function | ||||||
| Biological species |  METHANOCOCCUS THERMOLITHOTROPHICUS (archaea) METHANOCOCCUS THERMOLITHOTROPHICUS (archaea) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.8 Å MOLECULAR REPLACEMENT / Resolution: 2.8 Å | ||||||
 Authors Authors | Tishchenko, S.V. / Nevskaya, N.A. / Pavelyev, M.N. / Nikonov, S.V. / Garber, M.B. / Piendl, W. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 2002 Journal: Acta Crystallogr.,Sect.D / Year: 2002Title: Structure of Ribosomal Protein L1 from Methanococcus Thermolithotrophicus. Functionally Important Structural Invariants on the L1 Surface Authors: Nevskaya, N.A. / Tishchenko, S.V. / Paveliev, M. / Smolinskaya, Y. / Fedorov, R. / Piendl, W. / Nakamura, Y. / Toyoda, T. / Garber, M.B. / Nikonov, S.V. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1dwu.cif.gz 1dwu.cif.gz | 89.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1dwu.ent.gz pdb1dwu.ent.gz | 70.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1dwu.json.gz 1dwu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dw/1dwu https://data.pdbj.org/pub/pdb/validation_reports/dw/1dwu ftp://data.pdbj.org/pub/pdb/validation_reports/dw/1dwu ftp://data.pdbj.org/pub/pdb/validation_reports/dw/1dwu | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1cjsS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 23831.873 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  METHANOCOCCUS THERMOLITHOTROPHICUS (archaea) METHANOCOCCUS THERMOLITHOTROPHICUS (archaea)Strain: SN-1 / Description: GERMAN COLLECTION OF MICROORGANISMS (DSM) / Cellular location: RIBOSOME / Gene: THE CORRESPONDING GENE IN E. COLI IS RPLA / Plasmid: PMTHL1.4 / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.63 Å3/Da / Density % sol: 53 % | |||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 5.5 Details: 10% PEG 4K, 40MM MGCL2, 50MM NAAC PH 5.5, 50MM NACL, AGAINST: 30% PEG 4K, 100 MM NAAC | |||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 277 K / pH: 8.5 / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 |
| Detector | Type: R-AXIS IV / Detector: IMAGE PLATE / Details: MIRRORS |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.68→31.01 Å / Num. obs: 13590 / % possible obs: 95.1 % / Observed criterion σ(I): 0 / Redundancy: 4.31 % / Rmerge(I) obs: 0.072 |
| Reflection shell | Resolution: 2.68→2.77 Å / % possible all: 78.3 |
| Reflection | *PLUS Highest resolution: 2.7 Å / Lowest resolution: 30 Å / Num. obs: 14280 / % possible obs: 94.6 % / Rmerge(I) obs: 0.07 |
| Reflection shell | *PLUS % possible obs: 81.7 % / Redundancy: 3.8 % / Rmerge(I) obs: 0.275 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1CJS Resolution: 2.8→8 Å / Data cutoff high absF: 1000000 / Data cutoff low absF: 0.001 / Cross valid method: THROUGHOUT / σ(F): 2 / Details: X-PLOR V3.1F WAS ALSO USED.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 37.5 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.8→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.8→2.92 Å / Total num. of bins used: 8
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file | Serial no: 1 / Param file: PARAM19X.PRO / Topol file: TOPH19X.PRO | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.851 / Classification: refinement X-PLOR / Version: 3.851 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.7 Å / Lowest resolution: 30 Å / Num. reflection obs: 12624 / % reflection Rfree: 8.9 % / Rfactor Rfree: 0.254 / Rfactor Rwork: 0.189 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 53 Å2 |
 Movie
Movie Controller
Controller












 PDBj
PDBj

