+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1de1 | ||||||
|---|---|---|---|---|---|---|---|
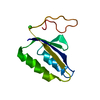
| Title | NMR STRUCTURES OF OXIDIZED BACTERIOPHAGE T4 GLUTAREDOXIN | ||||||
 Components Components | GLUTAREDOXIN | ||||||
 Keywords Keywords | ELECTRON TRANSPORT / GLUTAREDOXIN | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) | ||||||
| Method | SOLUTION NMR / SIMULATED ANNEALING, MOLECULAR DYNAMICS | ||||||
 Authors Authors | Wang, Y. / Amegbey, G. / Wishart, D.S. | ||||||
 Citation Citation |  Journal: J.Biomol.Nmr / Year: 2004 Journal: J.Biomol.Nmr / Year: 2004Title: Solution structures of reduced and oxidized bacteriophage T4 glutaredoxin. Authors: Wang, Y. / Amegbey, G. / Wishart, D.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1de1.cif.gz 1de1.cif.gz | 812.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1de1.ent.gz pdb1de1.ent.gz | 683.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1de1.json.gz 1de1.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/de/1de1 https://data.pdbj.org/pub/pdb/validation_reports/de/1de1 ftp://data.pdbj.org/pub/pdb/validation_reports/de/1de1 ftp://data.pdbj.org/pub/pdb/validation_reports/de/1de1 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 10063.631 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Details: THIS SEQUENCE OCCURS NATURALLY IN BACTERIOPHAGE T4 / Source: (natural)  Enterobacteria phage T4 (virus) / Genus: T4-like viruses / Species: Enterobacteria phage T4 sensu lato / References: UniProt: P00276 Enterobacteria phage T4 (virus) / Genus: T4-like viruses / Species: Enterobacteria phage T4 sensu lato / References: UniProt: P00276 |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|---|
| NMR experiment | Type: 2D NOESY, TOCSY, DQF COSY |
| NMR details | Text: THE STRUCTURES WERE DETERMINED USING STANDARD 2D HOMONUCLEAR TECHNIQUES |
- Sample preparation
Sample preparation
| Details | Contents: 2MM T4 GLUTAREDOXIN; 50MM PH7.0 PHOSPHATE BUFFER |
|---|---|
| Sample conditions | Ionic strength: 0 / pH: 7 / Pressure: AMBIENT / Temperature: 298 K |
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Varian UNITY / Manufacturer: Varian / Model: UNITY / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: SIMULATED ANNEALING, MOLECULAR DYNAMICS / Software ordinal: 1 Details: THE STRUCTURES ARE BASED ON 1050 DISTANCE, 188 DIHEDRAL ANGLE AND 201 H CHEMICAL SHIFT CONSTRAINTS. | ||||||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||||||
| NMR ensemble | Conformer selection criteria: BACK CALCULATED DATA AGREE WITH EXPERIMENTAL NOESY SPECTRUM,STRUCTURES WITH ACCEPTABLE COVALENT GEOMETRY,STRUCTURES WITH FAVORABLE NON-BOND ENERGY, STRUCTURES WITH THE ...Conformer selection criteria: BACK CALCULATED DATA AGREE WITH EXPERIMENTAL NOESY SPECTRUM,STRUCTURES WITH ACCEPTABLE COVALENT GEOMETRY,STRUCTURES WITH FAVORABLE NON-BOND ENERGY, STRUCTURES WITH THE LEAST RESTRAINT VIOLATIONS,STRUCTURES WITH THE LOWEST ENERGY,TARGET FUNCTION Conformers calculated total number: 50 / Conformers submitted total number: 30 |
 Movie
Movie Controller
Controller













 PDBj
PDBj X-PLOR
X-PLOR