+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1bqr | ||||||
|---|---|---|---|---|---|---|---|
| Title | REDUCED PSEUDOAZURIN | ||||||
 Components Components | PSEUDOAZURIN | ||||||
 Keywords Keywords | ELECTRON TRANSPORT / CUPROPROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Achromobacter cycloclastes (bacteria) Achromobacter cycloclastes (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.6 Å MOLECULAR REPLACEMENT / Resolution: 1.6 Å | ||||||
 Authors Authors | Inoue, T. / Nishio, N. / Hamanaka, S. / Shimomura, T. / Harada, S. / Suzuki, S. / Kohzuma, T. / Shidara, S. / Iwasaki, H. / Kai, Y. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 1999 Journal: J.Biol.Chem. / Year: 1999Title: Crystal structure determinations of oxidized and reduced pseudoazurins from Achromobacter cycloclastes. Concerted movement of copper site in redox forms with the rearrangement of hydrogen bond ...Title: Crystal structure determinations of oxidized and reduced pseudoazurins from Achromobacter cycloclastes. Concerted movement of copper site in redox forms with the rearrangement of hydrogen bond at a remote histidine. Authors: Inoue, T. / Nishio, N. / Suzuki, S. / Kataoka, K. / Kohzuma, T. / Kai, Y. #1:  Journal: Thesis Journal: ThesisTitle: Crystal Structure Determinations of Oxidized and Reduced Pseudoazurin from Achromobacter Cycloclastes: A Redox-Induced Conformational Change Contains a Peptide Bond Flip #2:  Journal: J.Biochem.(Tokyo) / Year: 1993 Journal: J.Biochem.(Tokyo) / Year: 1993Title: Crystallization and Preliminary X-Ray Studies on Pseudoazurin from Achromobacter Cycloclastes Iam1013 Authors: Inoue, T. / Nishio, N. / Kai, Y. / Harada, S. / Ohshiro, Y. / Suzuki, S. / Kohzuma, T. / Shidara, S. / Iwasaki, H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1bqr.cif.gz 1bqr.cif.gz | 36.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1bqr.ent.gz pdb1bqr.ent.gz | 24.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1bqr.json.gz 1bqr.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bq/1bqr https://data.pdbj.org/pub/pdb/validation_reports/bq/1bqr ftp://data.pdbj.org/pub/pdb/validation_reports/bq/1bqr ftp://data.pdbj.org/pub/pdb/validation_reports/bq/1bqr | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1bqkC  1pzaS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 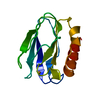
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 13032.940 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Details: REDUCED BY ASCORBIC ACID / Source: (natural)  Achromobacter cycloclastes (bacteria) / Strain: IAM1013 / References: UniProt: P19567 Achromobacter cycloclastes (bacteria) / Strain: IAM1013 / References: UniProt: P19567 |
|---|---|
| #2: Chemical | ChemComp-CU / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.9 Å3/Da / Density % sol: 40 % | |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 6 / Details: pH 6.0 | |||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / Method: vapor diffusion, hanging drop / Details: Inoue, T., (1993) J.Biochem.(Tokyo), 114, 761. | |||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 |
| Detector | Type: RIGAKU RAXIS IIC / Detector: IMAGE PLATE / Date: Oct 13, 1995 |
| Radiation | Monochromator: GRAPHITE(002) / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.6→30 Å / Num. obs: 13287 / % possible obs: 90.4 % / Observed criterion σ(I): 1 / Redundancy: 4.4 % / Biso Wilson estimate: 16.97 Å2 / Rmerge(I) obs: 0.044 / Net I/σ(I): 10.5 |
| Reflection shell | Resolution: 1.6→1.66 Å / Redundancy: 2.5 % / Rmerge(I) obs: 0.141 / % possible all: 70.8 |
| Reflection | *PLUS Num. measured all: 58870 |
| Reflection shell | *PLUS Highest resolution: 1.6 Å / % possible obs: 70.8 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1PZA Resolution: 1.6→8 Å / Cross valid method: THROUGHOUT Details: ESTIMATED COORDINATE ERROR. ESD FROM LUZZATI PLOT (A) : 0.153
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 18.2 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj


