+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1a7z | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CRYSTAL STRUCTURE OF ACTINOMYCIN Z3 | ||||||||||||
 Components Components | ACTINOMYCIN Z3 | ||||||||||||
 Keywords Keywords | ANTIBIOTIC / ACTINOMYCIN D / ACTINOMYCIN / ANTICANCER / ANTITUMOR / CHROMOPHORE / DEPSIPEPTIDE | ||||||||||||
| Function / homology | Actinomycin Z3 / BENZENE / :  Function and homology information Function and homology information | ||||||||||||
| Biological species |  STREPTOMYCES FRADIAE (bacteria) STREPTOMYCES FRADIAE (bacteria) | ||||||||||||
| Method |  X-RAY DIFFRACTION / AB INITIO / Resolution: 0.95 Å X-RAY DIFFRACTION / AB INITIO / Resolution: 0.95 Å | ||||||||||||
 Authors Authors | Schafer, M. | ||||||||||||
 Citation Citation |  Journal: Angew.Chem.Int.Ed.Engl. / Year: 1998 Journal: Angew.Chem.Int.Ed.Engl. / Year: 1998Title: Crystal Structures of Actinomycin D and Actinomycin Z3. Authors: Schafer, M. / Sheldrick, G.M. / Bahner, I. / Lackner, H. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1a7z.cif.gz 1a7z.cif.gz | 27.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1a7z.ent.gz pdb1a7z.ent.gz | 20.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1a7z.json.gz 1a7z.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/a7/1a7z https://data.pdbj.org/pub/pdb/validation_reports/a7/1a7z ftp://data.pdbj.org/pub/pdb/validation_reports/a7/1a7z ftp://data.pdbj.org/pub/pdb/validation_reports/a7/1a7z | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | 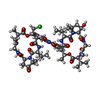  Type: Polypeptide / Class: Antibiotic / Mass: 1355.875 Da / Num. of mol.: 2 / Source method: isolated from a natural source Type: Polypeptide / Class: Antibiotic / Mass: 1355.875 Da / Num. of mol.: 2 / Source method: isolated from a natural sourceDetails: ACTINOMYCIN Z3 CONSISTS OF TWO PENTAMER RINGS LINKED BY THE CHROMOPHORE (PXZ) Source: (natural)  STREPTOMYCES FRADIAE (bacteria) / References: NOR: NOR00237, Actinomycin Z3 STREPTOMYCES FRADIAE (bacteria) / References: NOR: NOR00237, Actinomycin Z3#2: Chemical | ChemComp-BNZ / #3: Water | ChemComp-HOH / | Compound details | ACTINOMYCIN Z3 IS A BICYCLIC PEPTIDE, A MEMBER OF THE ACTINOMYCIN FAMILY. HERE, ACTINOMYCIN D IS ...ACTINOMYCI | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.4 Å3/Da / Density % sol: 49 % |
|---|---|
| Crystal grow | pH: 7 / Details: PH 7 |
| Crystal grow | *PLUS Method: unknown |
| Components of the solutions | *PLUS Common name: benzene |
-Data collection
| Diffraction | Mean temperature: 173 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: SIEMENS / Wavelength: 1.5418 ROTATING ANODE / Type: SIEMENS / Wavelength: 1.5418 |
| Detector | Type: SIEMENS HI-STAR / Detector: AREA DETECTOR / Date: Sep 22, 1997 / Details: COLLIMATOR |
| Radiation | Monochromator: GRAPHITE(002) / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 0.95→32.45 Å / Num. obs: 15653 / % possible obs: 99.5 % / Redundancy: 7.06 % / Rmerge(I) obs: 0.06 / Net I/σ(I): 8.17 |
| Reflection shell | Resolution: 0.95→1.05 Å / Redundancy: 4.85 % / Rmerge(I) obs: 0.35 / Mean I/σ(I) obs: 4.93 / % possible all: 98.1 |
| Reflection | *PLUS Num. measured all: 112696 / Rmerge(I) obs: 0.0602 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: AB INITIO / Resolution: 0.95→32.45 Å / Num. parameters: 3024 / Num. restraintsaints: 5470 / Cross valid method: FREE R / σ(F): 0 StereochEM target val spec case: NO RESTRAINTS ON DISTANCES AND ANGLES Stereochemistry target values: ENGH AND HUBER Details: ANISOTROPIC REFINEMENT REDUCED FREE R (NO CUTOFF) BY 0.026
| |||||||||||||||||||||||||||||||||
| Refine analyze | Num. disordered residues: 5 / Occupancy sum hydrogen: 286 / Occupancy sum non hydrogen: 307 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 0.95→32.45 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||
| Software | *PLUS Name: SHELXL / Version: 97 / Classification: refinement | |||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rwork: 0.0797 | |||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: s_angle_d |
 Movie
Movie Controller
Controller









 PDBj
PDBj