[English] 日本語
 Yorodumi
Yorodumi- EMDB-9945: Intact human mitochondrial calcium uniporter complex with MICU1/M... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9945 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Intact human mitochondrial calcium uniporter complex with MICU1/MICU2 subunits | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | ion channel / mitochondrial Ca2+ uptake / MCU / MICU / TRANSPORT PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationmitochondrial crista junction / negative regulation of mitochondrial calcium ion concentration / regulation of cellular hyperosmotic salinity response / positive regulation of cristae formation / uniporter activity / uniplex complex / Processing of SMDT1 / positive regulation of mitochondrial calcium ion concentration / Mitochondrial calcium ion transport / mitochondrial calcium ion transmembrane transport ...mitochondrial crista junction / negative regulation of mitochondrial calcium ion concentration / regulation of cellular hyperosmotic salinity response / positive regulation of cristae formation / uniporter activity / uniplex complex / Processing of SMDT1 / positive regulation of mitochondrial calcium ion concentration / Mitochondrial calcium ion transport / mitochondrial calcium ion transmembrane transport / mitochondrial calcium ion homeostasis / calcium import into the mitochondrion / calcium ion sensor activity / channel activator activity / calcium ion import / cellular response to calcium ion starvation / positive regulation of neutrophil chemotaxis / positive regulation of mitochondrial fission / protein complex oligomerization / calcium channel inhibitor activity / calcium channel complex / Mitochondrial protein degradation / cellular response to calcium ion / generation of precursor metabolites and energy / calcium channel regulator activity / calcium-mediated signaling / defense response / protein homooligomerization / mitochondrial intermembrane space / mitochondrial membrane / calcium channel activity / positive regulation of insulin secretion / cell migration / glucose homeostasis / protein-macromolecule adaptor activity / cell population proliferation / mitochondrial inner membrane / mitochondrial matrix / protein heterodimerization activity / calcium ion binding / mitochondrion / nucleoplasm / identical protein binding Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | ||||||||||||
 Authors Authors | Zhuo W / Zhou H | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Protein Cell / Year: 2021 Journal: Protein Cell / Year: 2021Title: Structure of intact human MCU supercomplex with the auxiliary MICU subunits. Authors: Wei Zhuo / Heng Zhou / Runyu Guo / Jingbo Yi / Laixing Zhang / Lei Yu / Yinqiang Sui / Wenwen Zeng / Peiyi Wang / Maojun Yang /  | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9945.map.gz emd_9945.map.gz | 12.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9945-v30.xml emd-9945-v30.xml emd-9945.xml emd-9945.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9945.png emd_9945.png | 103.5 KB | ||
| Filedesc metadata |  emd-9945.cif.gz emd-9945.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9945 http://ftp.pdbj.org/pub/emdb/structures/EMD-9945 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9945 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9945 | HTTPS FTP |
-Related structure data
| Related structure data |  6k7yMC  9944C  6k7xC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9945.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9945.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.091 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
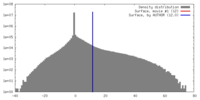
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Intact human mitochondrial calcium uniporter complex with MICU1/M...
| Entire | Name: Intact human mitochondrial calcium uniporter complex with MICU1/MICU2 subunits |
|---|---|
| Components |
|
-Supramolecule #1: Intact human mitochondrial calcium uniporter complex with MICU1/M...
| Supramolecule | Name: Intact human mitochondrial calcium uniporter complex with MICU1/MICU2 subunits type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Calcium uniporter protein, mitochondrial
| Macromolecule | Name: Calcium uniporter protein, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32.300203 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DDVTVVYQNG LPVISVRLPS RRERCQFTLK PISDSVGVFL RQLQEEDRGI DRVAIYSPDG VRVAASTGID LLLLDDFKLV INDLTYHVR PPKRDLLSHE NAATLNDVKT LVQQLYTTLC IEQHQLNKER ELIERLEDLK EQLAPLEKVR IEISRKAEKR T TLVLWGGL ...String: DDVTVVYQNG LPVISVRLPS RRERCQFTLK PISDSVGVFL RQLQEEDRGI DRVAIYSPDG VRVAASTGID LLLLDDFKLV INDLTYHVR PPKRDLLSHE NAATLNDVKT LVQQLYTTLC IEQHQLNKER ELIERLEDLK EQLAPLEKVR IEISRKAEKR T TLVLWGGL AYMATQFGIL ARLTWWEYSW DIMEPVTYFI TYGSAMAMYA YFVMTRQEYV YPEARDRQYL LFFHKGAKKS RF DLEKYNQ LKDAIAQAEM DLKRLRDPLQ VHLPLRQIG UniProtKB: Calcium uniporter protein, mitochondrial |
-Macromolecule #2: Essential MCU regulator, mitochondrial
| Macromolecule | Name: Essential MCU regulator, mitochondrial / type: protein_or_peptide / ID: 2 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 5.993192 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VIVTRSGAIL PKPVKMSFGL LRVFSIVIPF LYVGTLISKN FAALLEEHDI FVPE UniProtKB: Essential MCU regulator, mitochondrial |
-Macromolecule #3: Calcium uptake protein 1, mitochondrial
| Macromolecule | Name: Calcium uptake protein 1, mitochondrial / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 54.432258 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFRLNSLSAL AELAVGSRWY HGGSQPIQIR RRLMMVAFLG ASAVTASTGL LWKRAHAESP PCVDNLKSDI GDKGKNKDEG DVCNHEKKT ADLAPHPEEK KKKRSGFRDR KVMEYENRIR AYSTPDKIFR YFATLKVISE PGEAEVFMTP EDFVRSITPN E KQPEHLGL ...String: MFRLNSLSAL AELAVGSRWY HGGSQPIQIR RRLMMVAFLG ASAVTASTGL LWKRAHAESP PCVDNLKSDI GDKGKNKDEG DVCNHEKKT ADLAPHPEEK KKKRSGFRDR KVMEYENRIR AYSTPDKIFR YFATLKVISE PGEAEVFMTP EDFVRSITPN E KQPEHLGL DQYIIKRFDG KKISQEREKF ADEGSIFYTL GECGLISFSD YIFLTTVLST PQRNFEIAFK MFDLNGDGEV DM EEFEQVQ SIIRSQTSMG MRHRDRPTTG NTLKSGLCSA LTTYFFGADL KGKLTIKNFL EFQRKLQHDV LKLEFERHDP VDG RITERQ FGGMLLAYSG VQSKKLTAMQ RQLKKHFKEG KGLTFQEVEN FFTFLKNIND VDTALSFYHM AGASLDKVTM QQVA RTVAK VELSDHVCDV VFALFDCDGN GELSNKEFVS IMKQRLMRGL EKPKDMGFTR LMQAMWKCAQ ETAWDFALPK Q UniProtKB: Calcium uptake protein 1, mitochondrial |
-Macromolecule #4: Calcium uptake protein 2, mitochondrial
| Macromolecule | Name: Calcium uptake protein 2, mitochondrial / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.852695 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: PSLRKQRFMQ FSSLEHEGEY YMTPRDFLFS VMFEQMERKT SVKKLTKKDI EDTLSGIQTA GCGSTFFRDL GDKGLISYTE YLFLLTILT KPHSGFHVAF KMLDTDGNEM IEKREFFKLQ KIISKQDDLM TVKTNETGYQ EAIVKEPEIN TTLQMRFFGK R GQRKLHYK ...String: PSLRKQRFMQ FSSLEHEGEY YMTPRDFLFS VMFEQMERKT SVKKLTKKDI EDTLSGIQTA GCGSTFFRDL GDKGLISYTE YLFLLTILT KPHSGFHVAF KMLDTDGNEM IEKREFFKLQ KIISKQDDLM TVKTNETGYQ EAIVKEPEIN TTLQMRFFGK R GQRKLHYK EFRRFMENLQ TEIQEMEFLQ FSKGLSFMRK EDFAEWLLFF TNTENKDIYW KNVREKLSAG ESISLDEFKS FC HFTTHLE DFAIAMQMFS LAHRPVRLAE FKRAVKVATG QELSNNILDT VFKIFDLDGD ECLSHEEFLG VLKNRMHRGL WVP QHQSIQ EYWKCVKKES UniProtKB: Calcium uptake protein 2, mitochondrial |
-Macromolecule #5: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #6: CARDIOLIPIN
| Macromolecule | Name: CARDIOLIPIN / type: ligand / ID: 6 / Number of copies: 8 / Formula: CDL |
|---|---|
| Molecular weight | Theoretical: 1.464043 KDa |
| Chemical component information |  ChemComp-CDL: |
-Macromolecule #7: (9R,11S)-9-({[(1S)-1-HYDROXYHEXADECYL]OXY}METHYL)-2,2-DIMETHYL-5,...
| Macromolecule | Name: (9R,11S)-9-({[(1S)-1-HYDROXYHEXADECYL]OXY}METHYL)-2,2-DIMETHYL-5,7,10-TRIOXA-2LAMBDA~5~-AZA-6LAMBDA~5~-PHOSPHAOCTACOSANE-6,6,11-TRIOL type: ligand / ID: 7 / Number of copies: 16 / Formula: PLX |
|---|---|
| Molecular weight | Theoretical: 767.132 Da |
| Chemical component information |  ChemComp-PLX: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.0) / Number images used: 91728 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
-Atomic model buiding 1
| Initial model |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Refinement | Protocol: FLEXIBLE FIT | ||||||||
| Output model |  PDB-6k7y: |
 Movie
Movie Controller
Controller













 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)