+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9197 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mouse Protocadherin gamma B6 dimer-of-dimers | ||||||||||||||||||
 Map data Map data | subtomogram averaged map of Protocadherin gamma B6 EC1-6 dimer-of-dimers in solution | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Biological species |   | ||||||||||||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 35.0 Å | ||||||||||||||||||
 Authors Authors | Brasch J / Noble AJ / Shapiro L / Carragher B / Potter CS | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: Visualization of clustered protocadherin neuronal self-recognition complexes. Authors: Julia Brasch / Kerry M Goodman / Alex J Noble / Micah Rapp / Seetha Mannepalli / Fabiana Bahna / Venkata P Dandey / Tristan Bepler / Bonnie Berger / Tom Maniatis / Clinton S Potter / Bridget ...Authors: Julia Brasch / Kerry M Goodman / Alex J Noble / Micah Rapp / Seetha Mannepalli / Fabiana Bahna / Venkata P Dandey / Tristan Bepler / Bonnie Berger / Tom Maniatis / Clinton S Potter / Bridget Carragher / Barry Honig / Lawrence Shapiro /  Abstract: Neurite self-recognition and avoidance are fundamental properties of all nervous systems. These processes facilitate dendritic arborization, prevent formation of autapses and allow free interaction ...Neurite self-recognition and avoidance are fundamental properties of all nervous systems. These processes facilitate dendritic arborization, prevent formation of autapses and allow free interaction among non-self neurons. Avoidance among self neurites is mediated by stochastic cell-surface expression of combinations of about 60 isoforms of α-, β- and γ-clustered protocadherin that provide mammalian neurons with single-cell identities. Avoidance is observed between neurons that express identical protocadherin repertoires, and single-isoform differences are sufficient to prevent self-recognition. Protocadherins form isoform-promiscuous cis dimers and isoform-specific homophilic trans dimers. Although these interactions have previously been characterized in isolation, structures of full-length protocadherin ectodomains have not been determined, and how these two interfaces engage in self-recognition between neuronal surfaces remains unknown. Here we determine the molecular arrangement of full-length clustered protocadherin ectodomains in single-isoform self-recognition complexes, using X-ray crystallography and cryo-electron tomography. We determine the crystal structure of the clustered protocadherin γB4 ectodomain, which reveals a zipper-like lattice that is formed by alternating cis and trans interactions. Using cryo-electron tomography, we show that clustered protocadherin γB6 ectodomains tethered to liposomes spontaneously assemble into linear arrays at membrane contact sites, in a configuration that is consistent with the assembly observed in the crystal structure. These linear assemblies pack against each other as parallel arrays to form larger two-dimensional structures between membranes. Our results suggest that the formation of ordered linear assemblies by clustered protocadherins represents the initial self-recognition step in neuronal avoidance, and thus provide support for the isoform-mismatch chain-termination model of protocadherin-mediated self-recognition, which depends on these linear chains. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9197.map.gz emd_9197.map.gz | 4.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9197-v30.xml emd-9197-v30.xml emd-9197.xml emd-9197.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9197.png emd_9197.png | 15.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9197 http://ftp.pdbj.org/pub/emdb/structures/EMD-9197 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9197 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9197 | HTTPS FTP |
-Related structure data
| Related structure data |  9198C  9199C  9200C  6e6bC C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10234 (Title: Single particle cryoEM of mouse protocadherin gamma B6 EMPIAR-10234 (Title: Single particle cryoEM of mouse protocadherin gamma B6Data size: 438.8 / Data #1: Micrograph frames [micrographs - multiframe] Data #2: Micrographs along with all other magnification images from the collection [micrographs - single frame])  EMPIAR-10235 (Title: Single particle cryo electron tomography of mouse protocadherin gamma B6 EMPIAR-10235 (Title: Single particle cryo electron tomography of mouse protocadherin gamma B6Data size: 384.1 / Data #1: K2 tilt-series frames [micrographs - multiframe] Data #2: Whole-frame aligned tilt images along with all other magnification images from the collection [micrographs - single frame] Data #3: Appion-Protomo tilt-series alignments [tilt series]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_9197.map.gz / Format: CCP4 / Size: 5.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9197.map.gz / Format: CCP4 / Size: 5.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | subtomogram averaged map of Protocadherin gamma B6 EC1-6 dimer-of-dimers in solution | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
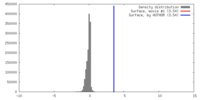
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : mouse protocadherin gamma b6
| Entire | Name: mouse protocadherin gamma b6 |
|---|---|
| Components |
|
-Supramolecule #1: mouse protocadherin gamma b6
| Supramolecule | Name: mouse protocadherin gamma b6 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: human embryonic kidney 293 freestyle / Recombinant plasmid: pi alpha Homo sapiens (human) / Recombinant cell: human embryonic kidney 293 freestyle / Recombinant plasmid: pi alpha |
| Molecular weight | Experimental: 300 KDa |
-Macromolecule #1: mouse protocadherin gamma b6
| Macromolecule | Name: mouse protocadherin gamma b6 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GQPVRYSIPE ELDRGSVVGK LAKDLGLSVL EVSSRKLRVS AEKLHFSVDS ESGDLLVKDR IDREQICKG RRKCELQLEA VLENPLNIFH VVVGIEDVND NAPQFEKKET RLEILETVAV G TRIPLEPA TDPDINLNSV KDYQISSNPY FSLMVRVNPD GGKTPELSLE ...String: GQPVRYSIPE ELDRGSVVGK LAKDLGLSVL EVSSRKLRVS AEKLHFSVDS ESGDLLVKDR IDREQICKG RRKCELQLEA VLENPLNIFH VVVGIEDVND NAPQFEKKET RLEILETVAV G TRIPLEPA TDPDINLNSV KDYQISSNPY FSLMVRVNPD GGKTPELSLE KLLDREEQRS HR LILTALD GGDPPRSATT QIEISVKDNN DNPPVFSKEE YWVSVSENLS PGSSVLQVTA TDE DEGVNA EILYYFRSTA QSTRHVFSLD EKTGVIKNNQ SLDFEDIERY TMEVEAKDGG GLST RCKII IEVLDENDNS PEITITSLSD HILENSPPGV VVVLFKTRDR DFGGNGEVTC DIGKD LPFK IQASSSNYYK LVTDGALDRE QNPQYNVTIT ATDKGKPALS SSTTIVLHIT DINDNA PAF QKSSYIVHVA ENNPPGASIA QVSASDPDLG ANGHVSYSII ASDLEPKSLW SYVTVNA QS GVVFAQRAFD HEQLRSFQLT LQARDQGKPS LSANVSMRVL VGDRNDNAPR VLYPALEP D GSALFDMVPR AAEPGYLVTK VVAVDADSGH NAWLSYHVLQ ASDPGLFSLG LRTGEVRTA RALGEKDAAR QRLLVGVRDG GQPPLSATAT LLLVFADSLQ EHHHHHHHH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.075 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||
| Grid | Material: COPPER / Support film - Material: CARBON / Support film - topology: LACEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: OTHER | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 298.15 K / Instrument: SPOTITON / Details: homemade nanowire grids. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 2.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Number classes used: 1 / Applied symmetry - Point group: C1 (asymmetric) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 35.0 Å / Resolution method: OTHER / Software - Name: Dynamo (ver. 109) / Details: FSC 0.143 map to model / Number subtomograms used: 506 |
|---|---|
| Extraction | Number tomograms: 7 / Number images used: 506 / Reference model: random subset / Method: volumes picked manually / Software - Name: Dynamo (ver. 1.1.109) / Software - details: Dipole picking Details: Volumes were picked using Dipole set models in Dynamo. Ends of the particles were marked using 'North' and 'South' options. |
| Final angle assignment | Type: OTHER / Software - Name: Dynamo (ver. 109) / Details: sub-volume correlation based alignment |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)