+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8984 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Icosahedral cryo-EM structure of mature Kunjin virus | |||||||||
 Map data Map data | Icosahedral reconstruction of mature Kunjin virus. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Kunjin virus (STRAIN MRM61C) Kunjin virus (STRAIN MRM61C) | |||||||||
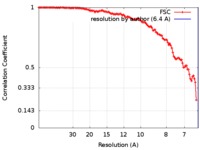
| Method | single particle reconstruction / cryo EM / Resolution: 6.4 Å | |||||||||
 Authors Authors | Therkelsen MD / Klose T / Vago F / Jiang W / Rossmann MG / Kuhn RJ | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2018 Journal: Proc Natl Acad Sci U S A / Year: 2018Title: Flaviviruses have imperfect icosahedral symmetry. Authors: Matthew D Therkelsen / Thomas Klose / Frank Vago / Wen Jiang / Michael G Rossmann / Richard J Kuhn /  Abstract: Flaviviruses assemble initially in an immature, noninfectious state and undergo extensive conformational rearrangements to generate mature virus. Previous cryo-electron microscopy (cryo-EM) ...Flaviviruses assemble initially in an immature, noninfectious state and undergo extensive conformational rearrangements to generate mature virus. Previous cryo-electron microscopy (cryo-EM) structural studies of flaviviruses assumed icosahedral symmetry and showed the concentric organization of the external glycoprotein shell, the lipid membrane, and the internal nucleocapsid core. We show here that when icosahedral symmetry constraints were excluded in calculating the cryo-EM reconstruction of an immature flavivirus, the nucleocapsid core was positioned asymmetrically with respect to the glycoprotein shell. The core was positioned closer to the lipid membrane at the proximal pole, and at the distal pole, the outer glycoprotein spikes and inner membrane leaflet were either perturbed or missing. In contrast, in the asymmetric reconstruction of a mature flavivirus, the core was positioned concentric with the glycoprotein shell. The deviations from icosahedral symmetry demonstrated that the core and glycoproteins have varied interactions, which likely promotes viral assembly and budding. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8984.map.gz emd_8984.map.gz | 97.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8984-v30.xml emd-8984-v30.xml emd-8984.xml emd-8984.xml | 10.8 KB 10.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8984_fsc.xml emd_8984_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_8984.png emd_8984.png | 165.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8984 http://ftp.pdbj.org/pub/emdb/structures/EMD-8984 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8984 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8984 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8984.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8984.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedral reconstruction of mature Kunjin virus. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.24 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
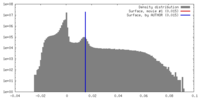
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Kunjin virus (STRAIN MRM61C)
| Entire | Name:  Kunjin virus (STRAIN MRM61C) Kunjin virus (STRAIN MRM61C) |
|---|---|
| Components |
|
-Supramolecule #1: Kunjin virus (STRAIN MRM61C)
| Supramolecule | Name: Kunjin virus (STRAIN MRM61C) / type: virus / ID: 1 / Parent: 0 / Details: Mature Kunjin virus purified from Vero cells. / NCBI-ID: 11078 / Sci species name: Kunjin virus (STRAIN MRM61C) / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Virus shell | Shell ID: 1 / Name: capsid shell / Diameter: 500.0 Å / T number (triangulation number): 3 |
-Supramolecule #2: Glycoprotein shell
| Supramolecule | Name: Glycoprotein shell / type: organelle_or_cellular_component / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Kunjin virus (STRAIN MRM61C) Kunjin virus (STRAIN MRM61C) |
-Supramolecule #3: Nucleocapsid core
| Supramolecule | Name: Nucleocapsid core / type: organelle_or_cellular_component / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Kunjin virus (STRAIN MRM61C) Kunjin virus (STRAIN MRM61C) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Material: COPPER / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Instrument: GATAN CRYOPLUNGE 3 / Details: Blot for 8 seconds before plunging.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 80.0 K / Max: 80.0 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 369 / Average electron dose: 24.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 18000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)