[English] 日本語
 Yorodumi
Yorodumi- EMDB-8293: Architecture of the Human Mitochondrial Iron-Sulfur Cluster Assem... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8293 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Architecture of the Human Mitochondrial Iron-Sulfur Cluster Assembly Machinery: the Complex Formed by the Iron Donor, the Sulfur Donor, and the Scaffold | |||||||||
 Map data Map data | Human Fe-S Assembly | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
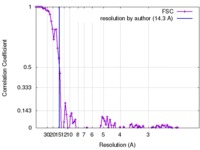
| Method | single particle reconstruction / negative staining / Resolution: 14.3 Å | |||||||||
 Authors Authors | Gakh O / Ranatunga W / Smith DY / Ahlgren EC / Al-Karadaghi S / Thompson JR / Isaya G | |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2016 Journal: J Biol Chem / Year: 2016Title: Architecture of the Human Mitochondrial Iron-Sulfur Cluster Assembly Machinery. Authors: Oleksandr Gakh / Wasantha Ranatunga / Douglas Y Smith / Eva-Christina Ahlgren / Salam Al-Karadaghi / James R Thompson / Grazia Isaya /   Abstract: Fe-S clusters, essential cofactors needed for the activity of many different enzymes, are assembled by conserved protein machineries inside bacteria and mitochondria. As the architecture of the human ...Fe-S clusters, essential cofactors needed for the activity of many different enzymes, are assembled by conserved protein machineries inside bacteria and mitochondria. As the architecture of the human machinery remains undefined, we co-expressed in Escherichia coli the following four proteins involved in the initial step of Fe-S cluster synthesis: FXN (iron donor); [NFS1]·[ISD11] (sulfur donor); and ISCU (scaffold upon which new clusters are assembled). We purified a stable, active complex consisting of all four proteins with 1:1:1:1 stoichiometry. Using negative staining transmission EM and single particle analysis, we obtained a three-dimensional model of the complex with ∼14 Å resolution. Molecular dynamics flexible fitting of protein structures docked into the EM map of the model revealed a [FXN]·[NFS1]·[ISD11]·[ISCU] complex, consistent with the measured 1:1:1:1 stoichiometry of its four components. The complex structure fulfills distance constraints obtained from chemical cross-linking of the complex at multiple recurring interfaces, involving hydrogen bonds, salt bridges, or hydrophobic interactions between conserved residues. The complex consists of a central roughly cubic [FXN]·[ISCU] sub-complex with one symmetric ISCU trimer bound on top of one symmetric FXN trimer at each of its eight vertices. Binding of 12 [NFS1]·[ISD11] sub-complexes to the surface results in a globular macromolecule with a diameter of ∼15 nm and creates 24 Fe-S cluster assembly centers. The organization of each center recapitulates a previously proposed conserved mechanism for sulfur donation from NFS1 to ISCU and reveals, for the first time, a path for iron donation from FXN to ISCU. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8293.map.gz emd_8293.map.gz | 18.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8293-v30.xml emd-8293-v30.xml emd-8293.xml emd-8293.xml | 11.1 KB 11.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8293_fsc.xml emd_8293_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_8293.png emd_8293.png | 85.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8293 http://ftp.pdbj.org/pub/emdb/structures/EMD-8293 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8293 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8293 | HTTPS FTP |
-Validation report
| Summary document |  emd_8293_validation.pdf.gz emd_8293_validation.pdf.gz | 79.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8293_full_validation.pdf.gz emd_8293_full_validation.pdf.gz | 78.2 KB | Display | |
| Data in XML |  emd_8293_validation.xml.gz emd_8293_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8293 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8293 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8293 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8293 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8293.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8293.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human Fe-S Assembly | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.034 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : NFS1-ISD11-ISCU-FXN
| Entire | Name: NFS1-ISD11-ISCU-FXN |
|---|---|
| Components |
|
-Supramolecule #1: NFS1-ISD11-ISCU-FXN
| Supramolecule | Name: NFS1-ISD11-ISCU-FXN / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism: |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Staining | Type: NEGATIVE / Material: 1% uranyl acetate / Details: 5 and 30 seconds | |||||||||
| Grid | Model: Carbon-coated copper grids, EMS / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Details: DV-502A vacuum evaporator (Denton Vacuum Inc.) | |||||||||
| Details | 50 mM Tris-HCl, pH 8.0, 150 mM NaCl |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 466 / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.0 µm / Calibrated defocus min: 0.21 µm / Calibrated magnification: 115000 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.21 µm / Nominal magnification: 115000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)